WEBLINE: A Novel Synthesized Immunotherapy Delivered Artificial Tertiary Layer Of The Immune System For Pan-Cancer Detection, & Treatment
Davyn Athwal
Delta West Academy
Grade 10
Presentation
No video provided
Problem
1.0 Abstract
The Scientific Initiative of WEBLINE (Widespread Epigenetically Budded Lymphoid-Iterative Neo-Lineage Endosystem) is one pertaining to the subject matter of sustained Pan Cancer remediation and remission, primarily addressing currently accepted clinical practice's inability to successfully treat and regularly eradicate Malignant Neoplasia. Typically, the vast majority of treatments and clinically applied practices that are utilized in today's world fail at even achieving a significant Overall Response Rate (ORR) without incurring an adverse reaction or creating unwanted byproducts of therapeutic intervention. WEBLINE aims to address this and many other issues that current clinical process's cause. Most particularly through the usage of genetically modified innate and adaptive immune cells which are modified to systematically neutralize Malignancy across a Pan Cancer spectrum through the use of currently existing immune processes and relative adaptability. Through this, a successful conclusionary result would be that of complete, cost efficient, risk free, and universally applicable Pan Cancer remediation through the usage of an artificial immune architecture to thoroughly and efficiently treat cancer.
2.0 An Introduction To Cancer
Cancer is an umbrella term describing over 200 different diseases causing widespread metastatic colonization leading to pleiotropic organo-axial dysfunction. Characterized by Pathogenic Genomic Aberrations Contributing To Constitutive Genomic Instability, metastasized malignancy induced Multiple Organ Dysfunction Syndrome (MODS), Hyperproliferative Neoplasia, and Widespread Immune Resistance, Cancer cells will rapidly mutate and replicate, taking crucial resources from healthy cells that require them for function [1], while forming tumors, breaking down healthy tissue, and spreading to other organs (metastasizing) which is currently the leading cause of all cancer related deaths [1]. Cancer is currently one of the leading killers in the world [1,2] with the conditions that this umbrella term pertains to claiming more than 9.7 million lives in 2022 alone. [2,3]. Currently treatments for cancer such as CAR-T Cell Therapy, Chemotherapy, CRISPR Cas9, Radiation therapy, hematopoietic stem cell transplantation (HSCT), Oncolytic Virotherapy, TIL therapy, and photo-based therapies have all represented significant advancements in oncological practices. [3,4,5] However, all of these therapies carry innate systemic toxicity, severe risk for patient tissue damage, Cytokine Rejection risk, Cytokine Release Syndrome risk, Exorbitant pecuniary burdens engendering critical allocative insufficiency which consequently imposes profound constraints upon the operational scalability and clinical applicability of these therapeutics in question, Non-selective Iatrogenic Histotoxicity resulting in the destruction and poisoning of healthy tissue, inherent operational inertia exacerbating malignant momentum, and the vast majority of therapeutics suffering from inability to combat broad scale Malignancies, being rigid and unadaptable against a foe that inherently adapts. Much of these therapeutics exhibit relatively low ORR's.
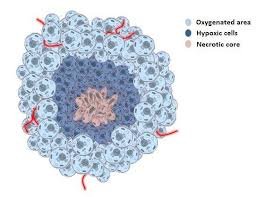
Fig 1: A Non Specified Diagram Pertaining To The Non Specified Structure Of A Solidified, Non Metastasized Malignancy.
2.1 Chimeric Antigen Receptor (CAR) T Cell Therapy
A clear example being Chimeric Antigen Receptor (CAR) T-cell therapy, despite lymphocytic cytotoxicity demonstrated by Chimeric Antigen Receptor (CAR) T-cell interventions against hematologic dyscrasias, these modalities exhibit a persistent therapeutic recalcitrance when directed toward desmoplastic solid-tumor microenvironments, such as ductal or lobular mammary carcinomas. Consequently, the application of extant protocols to these solidified malignancies fails to achieve sustained remission, instead precipitating a cascade of prohibitive fiscal expenditure, iatrogenic volatility, and a heightened propensity for severe adverse events, including cytokine release syndrome (CRS) and immune effector cell-associated neurotoxicity syndrome (ICANS). Not to mention the inherent temporal incubation period necessary for treatment application, CAR T Cell therapy cannot be administered without the necessary funds and lab conditions to construct the very cells, during which aggressive malignancies can and will progress further, and in some cases, become untreatable in contrast to how they were previously. [1,3,6,10].
2.2 Cytotoxic Pharmacotherapy (Chemotherapy)
Conventional antineoplastic pharmacotherapy such as the variations of Cytotoxic Chemotherapy are fundamentally constrained by a lack of biomolecular specificity, relying upon the indiscriminate disruption of mitotic kinetics which inevitably precipitates extensive collateral histopathological attrition. The systemic administration of these genotoxic agents necessitates a precarious titration between therapeutic efficacy and lethal systemic myelosuppression, frequently inducing iatrogenic multiorgan dysfunction and pro-inflammatory proteomic shifts. Furthermore, the inherent clonogenic heterogeneity and phenotypic plasticity of advanced malignancies render a universal 'pan-cancer' application effectively myopic; rather than achieving a targeted apoptotic resolution, these regimens often catalyze selective evolutionary pressure, facilitating the emergence of multidrug-resistant (MDR) subpopulations while simultaneously compromising the host’s immunological homeostasis. Furthermore, The aspiration for a universal antineoplastic paradigm via systemic chemotherapy is fundamentally obstructed by the stochastic nature of clonal evolution and the intrinsic molecular heterogeneity prevalent across divergent histotypes. Statistically, the therapeutic index for conventional cytotoxic regimens in late-stage solid-tumor malignancies remains disconcertingly narrow, with Phase III clinical trial attrition rates reaching as high as 90% [5]. In some meta-analyses. This pervasive failure is primarily attributed to apoptotic recalcitrance, wherein malignant subpopulations leverage the overexpression of anti-apoptotic BCL-2 family members and the downregulation of pro-apoptotic BH3-only proteins, effectively decoupling genotoxic stimuli from the caspase-mediated programmed cell death cascade. Furthermore, the systemic efficacy is 'diluted' by iatrogenic selection pressure, which inadvertently facilitates the expansion of chemo resistant subclones harboring ATP-binding cassette (ABC) transporter amplifications—such as P-glycoprotein (MDR1)—that orchestrate accelerated xenobiotic efflux. Consequently, rather than achieving a pan-cancer apoptotic resolution, these non-specific interventions often catalyze a transition toward refractory phenotypes and metastatic progression, underscoring a profound disconnect between controlled clinical trial efficacy and real-world oncological durability.
2.3 Hematopoietic Stem Cell Transplantation
The therapeutic trajectory of Hematopoietic Stem Cell Transplantation (HSCT) is frequently derailed by the fiscal enormity of the procedure—often exceeding $800,000 per allogeneic intervention—compounded by the profound risk of Graft-versus-Host Disease (GvHD). In vivo, the recipient undergoes a state of extreme immunological vulnerability and multiorgan attrition following myeloablative conditioning. This modality remains fundamentally monotypic, as it is incapable of addressing the polyclonal architecture of metastasized solid tumors, instead focusing narrowly on hematologic recovery while precipitating chronic iatrogenic endocrinopathies and a statistically significant shortening of the host’s biological lifespan. While not used to treat multiple Carcinomas directly, it is typically applied in tandem with Cytotoxic Pharmocotherapy, with the end goal being the regeneration of the patient’s bone marrow, however this only adds to the potential for complications and risks that pharmacotherapy causes inherently. Thus contributing further to overall risk of patient health deterioration as well as severely shortening life expectancy as said Stem cells exhaust at a largely expediated rate compared to that of naturally occurring bone marrow cells. Thus although not a treatment made to directly trigger a response or shrinkage of carcinomas, it still fails at it's current purpose of precipitating patient recovery and hematologic recovery allowing for temporary restoration of tissue that has been damaged as a result of currently accepted pharmacotherapies and carcinomas inherently. Thus adding yet another component that an active treatment must address, that component being patient recovery and tissue restoration conducted efficiently and risk free so as to not just eradicate a carcinoma, but allow for restoration of damage and return of the patient to full, natural functionality without that added intervention of further risky transplantation that can alter the patient in extremely negative manners.
2.4 Oncolytic Virotherapy and Tumor Infiltrating Lymphocyte Therapy
Oncolytic Virotherapy represents a step in the right direction for oncology, however it fails to negate the cons of pre existing carcinoma therapies and typically leads to severe side effects that severely hamper patient health or produce undesired effects. While Tumor-Infiltrating Lymphocyte (TIL) Therapy represents another right step, as it addresses and utilizes the immune system’s pre existing abilities to fight malignancies, however the side effects precipitated combined with the fiscal expenditure and overall therapy redundancy lead to a whole host of drawbacks. With manufacturing costs for personalized TIL products—such as Lifileucel—escalating toward $515,000 per cycle, rendering the treatment economically unsustainable for the general patient population. In vivo, viral vectors face rapid macrophage-mediated sequestration, preventing systemic reach, while TILs are rendered bioenergetically exhausted within the hypoxic, adenosine-rich tumor microenvironment (TME). These therapies exhibit a recalcitrance toward non-melanoma histotypes, failing to achieve pan-cancer durability while inducing severe vascular leak syndrome and pulmonary edema. While Oncolytic Virotherapies all typically share a tendency to be subjected to rapid mutation, effectively curating an infection within the body the hurts the innate tissue almost as much as the targeted malignancy, this combined with macrophage mediated sequestration leads to the vast majority of oncolytic virotherapies, although cheap, to either be nullified by the body’s internal defenses or rather damage the patient’s tissue further. Additionally, Tumor Infiltrating Lymphocyte’s have no way to restore energy or replenish ATP reserves as they are chronically starved for energy within the TME, this is further exacerbated by many malignancies often exhibiting Mitochondrial theft, causing TIL therapy to actually exacerbate and amplify the malignancies progression as each Lymphocyte acts as a massive energy store to any cancer cells that wish to extract the ATP. However notably, immunotherapies such as TIL therapy have shown promise and success against varying non solid malignancies, with as high as a 95% remission rate, as displayed in the data charts below. However, immunotherapies and general Carcinoma remediating therapies all struggle to achieve acceptable results opposite solid carcinomas. [2.8]
2.5 CRISPR-Cas9 Enzyme Therapy
The application of CRISPR-Cas9 as a systemic antineoplastic agent is currently obstructed by a prohibitive cost-per-patient ratio, reaching into the millions ($1M–$2M) for clinical-grade delivery. The primary in vivo flaw is the stochastic propensity for off-target mutagenesis, wherein unintended genomic rearrangements catalyze secondary oncogenesis (new malignancies). Furthermore, the immunogenicity of the Cas9 protein triggers a massive host-defense response, neutralizing the therapeutic agent before it can address the spatial heterogeneity of multiple metastatic sites, effectively rendering it an impotent modality for late-stage, broad-spectrum disease. Additionally, CRISPR-Cas9 is not an efficiently proper method of malignancy treatment, as it maintains a suboptimal-complete failure rate of cost to treatment impact ratio. As it by far pertains the largest fiscal burden of any malignancy treatment while simultaneously being relatively uncapable at dealing with many broad scale malignancies without large temporal expenditure for preparation not to mention the fiscal factors and the propensity for secondary malignancies being amplified to a grossly exorbitant rate.
2.6 Ionizing Radiation Therapy and Various Photodynamic Therapies
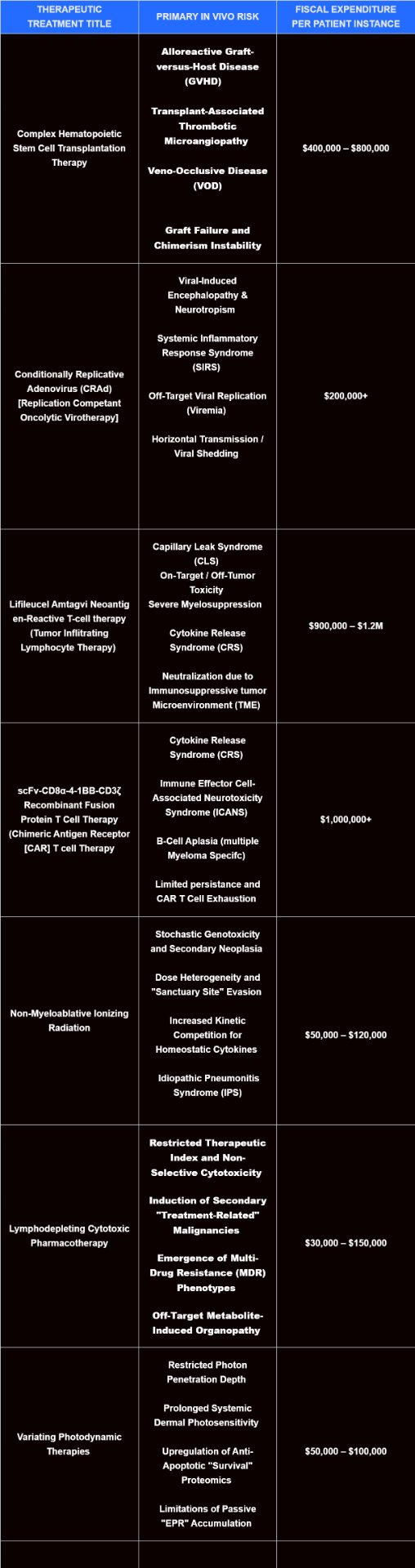
Ionizing Radiation (IR) and Photodynamic Therapy (PDT) suffer from spatiotemporal myopia, where the inability to address occult micrometastases renders them ineffective as pan-cancer solutions. IR frequently induces a 'badscopal' effect—an iatrogenic upregulation of the amphiregulin-CD47 axis—which paradoxically fuels the growth of distant, untreated tumors by creating a systemic immunosuppressive state. Photo-based therapies are further limited by negligible photon penetration in deep visceral tissues, making them biologically inert for internal solid tumors. The cumulative financial burden of long-term radiotherapy, combined with the risk of irremediable radiation-induced fibrosis, far outweighs the localized cytoreductive benefits in a disseminated disease state. Ionizing Radiation and variations of Photodynamic Therapy often are somewhat cost efficient, yet they have a large propensity and risk factor to deal more systemic tissue and organ damage and toxicity than a lower stage malignancy will itself. Additionally, photo-based therapies very frequently suffer from being unable to combat malignancies that are below the lower epidermis and located as deep as the vascular tissue. Leading to an inability to treat any metastasized solid malignancy or really any malignancy beyond epidermal malignancies such as acute Melanoma. Below we can see the costs and primary in vivo risk that is presented by currently existing treatments to cancer:
Fig 2: Simple Data Chart Describing Commonly Accepted Therapeutics & The Associated Risks and Fiscal Expenditure
2.7 The Necessity Of Safe, Cost Efficient, And Capably Adaptable Pan Cancer Remediation
As a result of these plentiful redundancies and issues pertaining to currently active and accepted medical treatments. The necessity to develop a therapy that is capable of pan cancer treatment on a broad scale is paramount to development of the medical industry’s capability to save lives. Cancer takes the lives of more than 9.7 million people per year [1]. While placing a global financial burden on the medical industry, siphoning almost 25.7 trillion dollars globally in treatments alone [6]. As such a vital therapeutic is of paramount necessity. Additionally, not just an efficient pan cancer therapeutic, but one that is cost efficient to allow for distribution in all areas of the world to all economic backgrounds, zero systemic toxicity to allow for risk free therapeutic application to patients afflicted with varying malignancies, while lastly but most certainly not least, a therapeutic that can properly address all the conditions that fall underneath the cancer umbrella term as opposed to just targeting one specific niche variation of malignancy. This way the treatment can be universally applied to all cancer patients, below we can see the pros and cons of currently available treatments, as well as the unfortunately high mortality rate for metastasized malignancies as it is readily known that late stage tumors can seldom be treated and even further seldom have a full recovery made. Below are charts, tables, and diagrams showcasing the remission rate and success aforementioned therapies have had in comparison to the cancer mortality rate as well as the in vivo complications that arise with the fiscal burden.
2.8 Speculative Research and Treatment Formulation: The Foundation For A Pan Cancer Solution
Currently, speculative research, clinical trials, in vivo treatments, and experimental application suggest that immunotherapies remain the most promising for a potential pan cancer solution. As our body, although ill equipped to properly address severely aggressive malignancies, contains the building blocks and the foundational principles that allow for malignancy treatment and utter eradication. Immunotherapies seek to enhance the immune system or deliver external components which assist or direct the immune system or externally inserted cells to attack and break apart cancerous tumors and assorted malignancies. The most prominent of which are TIL therapy (Tumor Infiltrating Lymphocyte Therapy) and Chimeric Antigen T Cell Therapy (CAR T Cell therapy). Both of which have had varying success. CAR T cell therapy excels at dispatching with liquid malignancies, most specifically assorted lymphomas and leukemias. However CAR T Cell therapy struggles with it’s tendency to cause extreme cytokine release syndrome as well as it’s tendency to fail due to hostile immune rejections and assorted autoimmune conditions either limiting or entirely nullifying the treatment’s capability or causing extensive further damage to the patient’s tissue and organ systems. While additionally being vastly limited by fiscal strains, with CAR T cell therapy even in dosages as small as 100,000 cells costing at the bare minimum for acquisition being $400,000+ per clinical instance. This figure only increases with the number of CAR T cells inserted into the patient and the severity of the malignance which in turn exacerbates the innate systemic risks that immunotherapies all carry. Whereas Tumor Infiltrating Lymphocyte therapy is a step in the right direction once again, with the therapy integrally being made up of cells that are preexisting within the body, cells that have proven they can infiltrate the TME and begin to kill cancer cells. Although this carries intense systemic risks, and furthermore the modifications to the lymphocytes are much too minor, not circumventing the innate hostility that the TME possesses. As innate Lymphocytes are often deactivated or killed through specific enzyme release by the TME, while additionally the metabolic wasteland that the is the TME causes any cells that make it past the initial wave to perish of hunger and starvation. Most immunotherapies are purely speculative right now due to their severe drawbacks. However, because of the success of previous immunotherapies and artificial immune architectures with regards to cancer treatment, it is ideal and most optimal to create an artificial immune architecture delivered via immunotherapy for pan cancer eradication. Therefore, ideally repurposing the Immune System’s innate bacterial and viral defenses into a complex malignancy treatment system. Below, we can observe the remission chance and ability of current treatments:
2.9 A Pan Cancer Solution: The Foundational Premise Of a Widespread Epigenetically-Budded Lymphoid-Iterative Neo-lineage Endosystem (WEBLINE) Immunotherapy
As we can see, Most immunotherapies have the correct foundational idea, however they lack the correct execution and mechanisms to formulate a risk free and optimal treatment, for this reason I aim to create and synthesize an immunotherapy that functions as a full pan cancer, risk free, remission proven, and fully adaptable artificial tertiary layer of the immune system delivered immunotherapy that is capable of consistently high Pan Cancer ORR and CR's, in contrast to currently accepted treatments which are rigid and volatile by nature. The immune system is inherently capable of malignancy eradication, only beginning to struggle against large, metastasized, solidified, and actively disarming TME’s and malignancies that deliberately weaken and hamper the body’s innate defenses. This is out of the lack of capability and lack of tools necessary to naturally eradicate this type of cellular foe in vivo, as the immune system has anti carcinomic functions as a subsidiary objective, as the primary directive of the immune system is to neutralize viral and bacterial incursion within the biological systems of the body. However, the main underlying functions that the immune system utilizes to combat infectious pathogens can be repurposed through genetic modification and neo-plasma mapping to instead primarily target carcinomas [1]. An example of this are Natural Killer T Cells. Which naturally trigger cellular apoptosis through protein release, these proteins bind to the receptors on cells and trigger their apoptosis cellular path. Initiating risk free and controlled cell death, by repurposing various innate and adaptive immune cells to utilize their functions to target malignancy through receptor, genetic, and directive variation and editing, a small number of cells capable of adapting to and neutralizing various different Carcinomas may be created, a naturally occurring solution that needn’t require the extensive fiscal, biological, and logistical issues that multiple artificial treatments face today. Firstly, the utilization and repurposing of immune cells in a low cost manner to act as a pan cancer remediation therapy would be paramount. Firstly being able to universally apply a therapy to any varied type of differentiating carcinoma would almost immediately remove many logistical issues that hospitals and doctors face today. While additionally having consistently high remission and overall response rates would lead to numerous more lives being saved as a result of treatment. While the inherent nature of the immune system is one of minimally altered defense, meaning rather than utilizing invasive and systemically hazardous and toxic treatments that inherently carry the risk of eliciting undesired and volatile outcomes, we could rather treat varied carcinomas utilizing a single therapy that due to it's inherent naturality, would be able to execute the desired outcome with minimal risk and minimal cost. Immunotherapies are currently a step in the right direction ontologically, it is just a matter of negating the risks and minimizing the costs while guaranteeing remission and carcinoma eradication while being able to allow a patient to be resistant to carcinomic incursion for the rest of their life. All goals that a Widespread Epigenetically Budded Lymphoid Iterative Neo Lineage Endo System of genetically modified and altered immune cells functioning as an artificially synthesized tertiary layer of the immune system could achieve, as to tackle a complex hostile Cellular Architecture such as a solidified, metastasized, late stage carcinoma, it is almost certainly necessary to possess a biological endo system or immune architecture of matching complexity to combat it. Thus by repurposing the immune system, one's very own inherent biological inhibitors to cellular foes to instead combat carcinomas as a complex immune architecture, while doing so in a cost efficient and risk free manner. A Pan Cancer Solution, what was once thought a myth, becomes a universally needed reality.
Method
3.0 Methodology For Therapeutic Formulation & Architectural Assembly
In order to formulate a universally applicable, cost efficient, Innocuous, and maximally efficacious therapeutic, we must first define the methodology and practice for which we will formulate, and test therapeutic efficacy. As we must first set parameters before moving forwards with formulation.
Materials
1. Literature Dive: Cases Provided Via Depersonalized Records Within The National Library Of Medicine\, Viewed To Establish Understanding In Core Disease [P] (For further information ask to see my Logbook/Therapeutic Formulation Document)
2. Software Acquisition: Benchling Programming Software\, Swissdock Molecular Docking Software & Protein Building\, Physicell Python Set\, Kbase Mathematical Database
3. Reference Various Articles Throughout (See Citations) To Ensure Accuracy & Biologically Sound Basis To Support Assertions With Regards To Therapeutic Advancement & Proper Analysis Of Informatics.
4. Utilized Each Of These Non Specific Cases As Testing Parameters For Therapeutic. Excluding One Case (Control) To Allow For Comparison With Regards To Clinically Accepted Practices
Anatomization Within Parameters Of Benchling
1. Constructed And Formulated New Project Complete With Folders and Sub Categories For WEBLINE Project
2. Utilized Literature Analysis to Formulate Understanding of Necessary Adenine\, Guanine\, Cyanine\, Thymine Bonds To Allow For Enabling Of Genetic Structure To Allow For Behavior & Cellular Capabilities to Be Altered To Elicit The Desired Effect\, Which Operate As The Building Blocks Of The Therapeutic's Inherent Function. Furthermore\, Upon Uploading Of Genetic AGCD Sequencing\, Utilized Benchling PRIMER75 & ANNOTATION6a Functions to Ensure Preciseness & Mark Cellular Components For Effect. While Allowing Benchling's Inherent Features To Artificially Construct Plasmid.
3. Performed Assurance Of Quality Tests Through The Utilization Of Uploading Datasets Pertaining To Other Currently Active Cells [Eg: Macrophage Antigen Presentation Circuit] In Order To Assure Software Capability To Accurately Formulate Plasmid.
4. Ensured Genetic Functions Within Differentiating Cells Complemented One Another With Maximal Therapeutic Efficacy
5. Preloaded PNGSA\, SNGSA\, & TNGSA Sequencing Groups Within Benchling\, Before Beginning Analysis for Penultimate Complementation Analysis Between Plasmids.
Primary Nucleic Genetic Sequencing Array (PNGSA)
First and Foremost genetic sequencing plasmid inserted into WEBLINE cell, functions to prevent autoimmune diseases and houses critical function within cell, with nucleic acids marked with AL5-09Alphf preventing somatic hypermutation within these genes. As these pertain to the prevention of autoimmune disease and furthermore, govern cellular quiescence (LuxS Enzymatic Response Gene) which governs an extremely important component with regards to the overarching therapeutic course.
Necessary as through the genetic effects, these are the levels of damage that autoimmune of histotoxic implications could do to a potential patient:
Where HALE = Healthy Applicable Life Expectancy, & R= Reduction due to therapeutics, and where Hp = Potential HALE, the following can be utilized:
Hp - R = HALER/HALE Reduction
| HALE Reduction By 1-5 Years | 21% of all Malignancy Survivors |
|---|---|
| HALE Reduction By 5-10 Years | 27% of all Malignancy Survivors |
| HALE Reduction By 10-20 Years | 30% of all Malignancy Survivors |
| HALE Reduction By 20+ Years | 11% of all Malignancy Survivors |
| Total Percentage of HALE Reduction among Malignancy Survivors | 89% of all |
| 11% remain with intact HALE. |
The above dataset illustrates the adverse effect that Malignant Neoplasia does to it's survivors even after full treatment. Either through failure to correctly restore systematic disassembly that has been inflicted upon the patient. Much of these HALE reductions are derived from treatment toxicity and issues regarding treatment application. Further emphasizing the necessity of both a new alternative and the necessity of the PNGSA, to prevent accidental Histotoxicity or indiscriminate cytotoxicity.
Secondary Nucleic Genetic Sequencing Array (SNGSA)
Secondary genetic sequencing plasmid inserted, functions with regards to therapeutic interactions with malignancy. Nucleic acids within this array are not marked with AL5-09Alphf which allow for protein induced regulated & manipulated somatic hypermutation to occur, permitting the altering of the inherent genetic code to formulate a genetic peptide reciprocal, functioning as the therapeutic's main method of adaptability and reactivation. Uploaded via Benchling as is with PNGSA.
Tertiary Nucleic Genetic Sequencing Array (TNGSA)
Functions to control and govern somatic hypermutation within each cell archetype. Whether between Macrophage, Neutrophil, Or Plasma cell variation. Each WEBLINE cell functions to allow for somatic hypermutation to permit adaptability to formulate a genetic reciprocal to what is developed upon the Malignancy to allow for optimal binding and the injection of the TNMA-5 Protein as detailed below.
These categorical plasmid arrays allowed me to break down the otherwise insurmountably complex cellular procedures and editing that I would need to undertake to procure the computational building blocks of the therapeutic in question. Furthermore, with these initial developments in Benchling I was able to computationally model and in silico produce the therapeutic in question, through the usage of the Complementation Analysis Through both Kbase & Benchling, I was able to make certain edits to various Arrays as needed, ensuring the therapeutic was capable of synergizing and performing in as an endosystem. Due to the nature of this being a novel therapeutic proposing a paradigm shifting approach towards pan cancer oncology, I must be sure that the actual principle behind the therapeutic in question is scientifically and biologically sound.
6. Allowed for Complementation Analysis to undergo\, which produced the SynPoten Statistic of 0.9984 pertaining to a 99.84% synergy towards the desired output effect\, cementing the capability of the plasmid insertions to elicit the cellular reaction and grant the means of biological malignancy eradication that I have designated for this therapeutic. For context:
| Cytotoxic Pharmacotherapy | 0.689 | 68.9% |
|---|---|---|
| Chimeric Antigen Receptor T Cell Therapy | 0.78 | 78% |
| WEBLINE | 0.9984 | 99.84% |
Fig 4: Table Displaying Results Derived From Similar Multi Omic Complementation Analysis From Another Genetic Therapeutic Mixed With An Analysis of Cytotoxic Pharmacotherapy Binding Affinity And Effect, Compartmentalized Into Table To Allow For Common Comparison Across A Mutual Medium
This showcases inherently already promising results from the analysis out of Benchling, Swissdock, & Kbase.
7. Utilized Seed Base Failure analysis to interpret how each genetic sequence within each of the proto WEBLINE cells may have failed\, through sequestering of each genetic circuit and subsequent analysis and re editing\, I was able to achieve the above pictured result\, thus concluding phase 1 of therapeutic development.
8. Initialized Swissdock For Protein building and receptor binding to orchestrate various planned therapeutic mechansims as depicted within the genetic circuit.
9. Viewed And Inferred Conclusionary Results Of Computational Analysis &
3.2 Benchling
Benchling is a plasmid mapping software that permits researchers in both the academic and pharmaceutical departments to engineer plasmids to illustrate components of gene editing or cellular biology. This software serves to allow for computational modeling of plasmid insertions to allow for the therapeutic to take full and official course. As WEBLINE itself is an immunotherapeutic necessitating the utilization of genetically modified cells operating in tandem, it is necessary to obtain a platform capable of modeling the genetic circuits, components, and elements required as prerequisites in order to facilitate the cellular capabilities and behaviors that we wish the therapeutic to perform under. As basic cellular modifications and genetic sequencing is the only governing factor differentiating a Plasma cell from producing polyclonal vs monoclonal antibodies, As such, in order to first orchestrate the building blocks of an immune architecture built upon genetically modified cells functioning in tandem as an endosystem, we must first utilize a computational modeling program within which we can define various genetic circuits and provide the plasmid insertions that are necessary for the modification of the immune cells in question. As my therapy hinges on the utilization of genetically modified cells, we must first develop a modicum to accurately and precisely define each genetic change/addition in order to formulate the therapeutic in question.
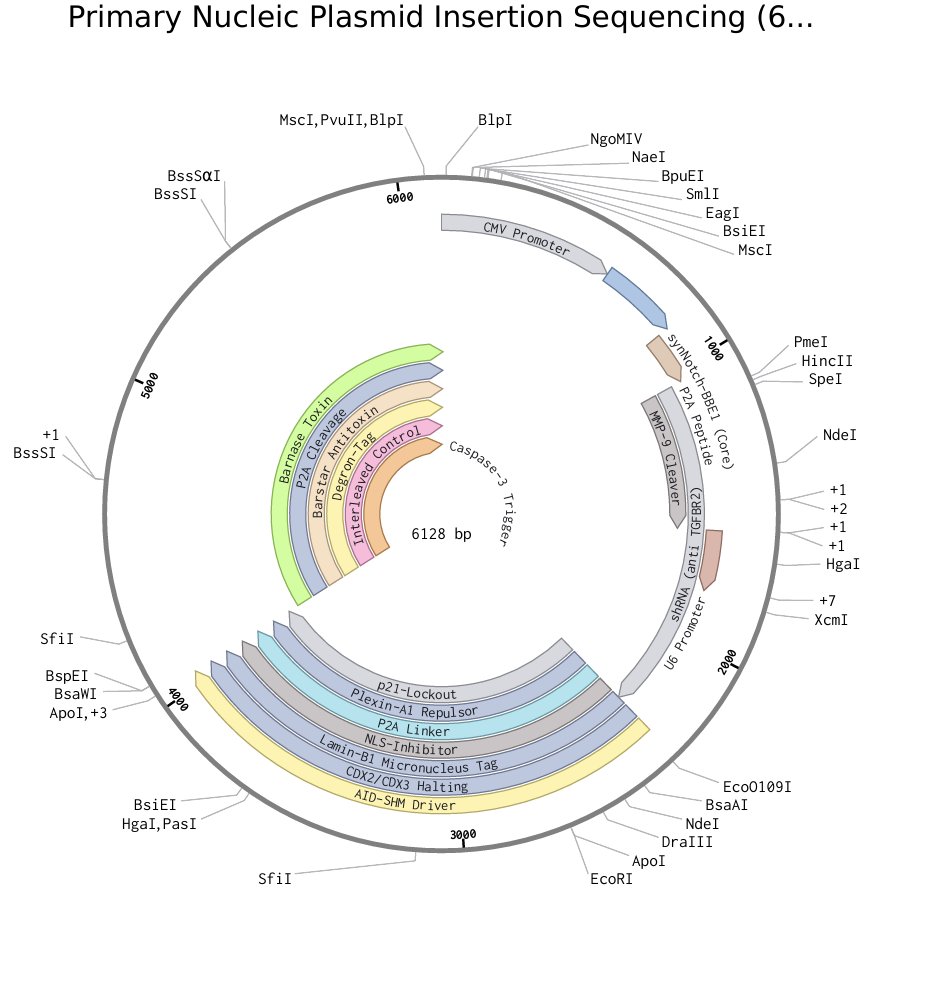 Fig 3: Example PNGSA Engineered Through Benchling, Purposed specifically For Epithelial Cell Variation, utilized specifically for genetic modification within extracted cells described in I-Phase (See Logbook)
Fig 3: Example PNGSA Engineered Through Benchling, Purposed specifically For Epithelial Cell Variation, utilized specifically for genetic modification within extracted cells described in I-Phase (See Logbook)
Fig 4: An Illustrative example of a Physicell Dataset Utilized Within The Parameters of My Trials, The Figure Above Displays 2 Day And 12 Hours In The Elapsed Simulation. Depicting The Length At Which Observations Occured
3.3 Swissdock
As the majority of WEBLINE cells initiate cellular apoptosis through the utilization of various receptors and antigen presentation, we must define the individual ability of receptors and components to bind with one another. Various protein modeling additionally needs to be formulated through the utilization of Swissdock, as various toxins and receptor interactions require efficient modeling to represent the therapeutic efficacy that we wish to achieve. This included but is not limited to:
-Receptor binding -Antibody Binding Capability -Toxin Injection & Chemical Composition -Overall Protein Modeling & Optimal Binding Potential
Furthermore, we must define the parameters and binding capability that these WEBLINE cells can exercise upon exposure to Malignant cells. As well as illustrate the structural and chemical properties of multiple molecular agents and various micro-organisms. Below we see the formation of two proteins and receptors which function as the WEBLINE cell's main methodology of triggering regulated cellular apoptosis.
Fig 5: Optimal Binding Position and Chemical Sequencing for Completed TNMA-1 Protein, Later To be Placed Upon TNMA-5 Monoclonal Antibody and To Be Presented Upon MTD-20 Receptor, As Well As On TNMA-5 Monoclonal Antibodies Secreted via Plasma Cell Apoptotic Variation-N.
Fig 6: Optimal Binding Position and Chemical Sequencing Model Of p53-PRRMA Protein, for presentation on Monoclonal Antibodies Within The Parameters Of The Plasma Cell Apoptotic Varation-R
3.4 Physicell
A program allowing for the representation of inherent immune cells capability to combat a large scale tumor through the utilization of python based coding. Within Physicell we can define rules based upon cellular behavior and cellular capability with regards to cytokine function, these rules are theoretically applicable within an in vitro and in vivo environment as the rule's practical functionality can be defined by inherent biological capabilities that each cellular structure within the therapeutic will possess. This program is utilized in order to prove therapeutic efficacy inferred through the application of an in silico modeling software to allow for actual therapeutic efficacy to be underneath analysis throughout the duration of testing procedures.
3.5 Kbase
Kbase is a program that allows for the metabolic modeling of a hypothetical cell through the usage of complex mathematical formula and defining of proper genetic structures and functions to allow for an accurate depiction of whether or not a unicellular organism can readily support itself from a metabolic standpoint. Furthermore we can model the hypothetical metabolic strain that an individual may be placed under throughout the duration of the therapeutic's efficacy. This way we can measure one of the most significant risks involved within any level of immunotherapeutic upon application to the human body.
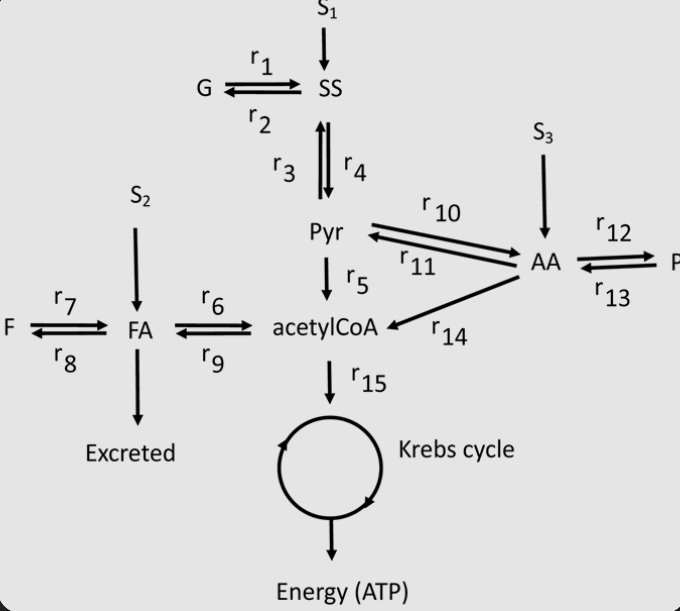 Fig 7: An Instance Of Kbase Metabolic Mapping Showcasing Carbon Metabolism, A Critical Component In Cellular ATP Production and Maintaining Metabolic Flow
Fig 7: An Instance Of Kbase Metabolic Mapping Showcasing Carbon Metabolism, A Critical Component In Cellular ATP Production and Maintaining Metabolic Flow
Fig 8: Formula Utilized For Metabolic Calculations With Regard To Epigenetically Edited Unicellular Organisms & For Overall Anatomic Metabolic Functioning With Inputted Components
3.6 Rationale & Computational Priming
My Rational for the utilization of these various programming pertains to the lack of currently available testing parameters. There is no perfect program to utilize for testing of a therapeutic bearing a magnitude such of that as WEBLINE. As a result, we must utilize Multi-Omic Analysis as depicted below:
Multi-Omic Assay & Deconstruction:
Due to the nature of this project, one must leverage the utility of multiple differentiating programs in order to facilitate and precipitate trials that hold impact with regard to the therapeutic in question. Firstly, Multi-Omic analysis is necessary as per my rational due to the fact that since no computational modeling software is capable of encompassing all the aspects of WEBLINE, we must utilize a multitude of computational software in order to precipitate both the maximal efficacy & Therapeutic comparison as illustrated prior.
1. Firstly\, therapeutic components and elements are constructed within Benchling/Kbase/Swissdock. Thus curating necessary elements of the aforementioned therapeutic. Through Multi-Omic Construction of these various elements\, such as the DNA and RNA structure of the plasmids for use within the cell as detailed within the therapeutic course. We can fabricate the foundational premise and building blocks of the therapeutic\, necessary through my rationale as these are necessary to justify the later stages of computational modeling and in silico validation.
2. Many of the mechanisms and molecular level interactions are facilitated through the usage of proteins which WEBLINE cells are given the capability of producing and leveraging. However it is of the utmost significance that we model these molecularly\, predicting the optimal bonding position of receptors\, binding affinity\, and define protein structure and chemical structure in order to completely ascertain that these specified proteins and molecular agents can successfully and efficaciously precipitate the denouement & conclusionary outcome that I wish of it within the parameters of WEBLINE. Ensuring that the Biological and Treatment stipulations that have been placed through the goals this project wishes to address are met through the use of complete testing. Furthermore\, we must leverage the usage of assurance of quality trials prior. As we must ensure the total punctiliousness and rigorous fastidiousness of the trials in question. As Biology is a precise endeavor\, we must also be precise.
3. Lastly I capitalized on the utility of Physicell\, a primarily C++ and Python framework\, acting as a base foundation for treatment modeling and therapeutic efficacy among other statistics. However\, due to it's nature as a primarily C++ framework\, this allows curators of testing that have downloaded the Physicell framework extreme creativity and customizing oppurtunity to tailor make their Physicell application to their needs with regards to in silico modeling. Moreover\, I harnessed the usage of Physicell by artificially modifying the program within a novel manner utilizing C++ coding and Python in accordance to allow for me to not only measure the therapeutic efficacy and statistics of WEBLINE\, but additionally to compare and contrast these with those of Cytotoxic Pharmacotherapy\, Chimeric Antigen Receptor T Cell Therapy\, VIRAS-9 Virotherapy\, Tumor Infiltrating Lymphocyte Therapy (TIL)\, Ionizing Radiation Therapy\, Crispr Cas9 Enzyme CLINIC Therapy (CC9CLINIC)\, and Variating Photodynamic Therapy (VPT). My Rationale being that this allowed me to properly and methodically analyze not just WEBLINE's efficacy with regards to malignant neoplasia\, but additionally allowed me to cross reference this with how well it performed in relation to already clinically accepted alternative treatments\, those of which I am trying to disprove. My rationale for utilizing Physicell however pertains to being able to prove the efficacy and statistical properties of WEBLINE in of itself\, as Physicell\, particularly with the addition of my own C++ blocks\, allows for the precise ORR of variating immune cells in response to both the TME (Tumor microenvironment) and the tumor in question. This also allowed me to further define the environmental and malignancy properties that I would be testing WEBLINE against. (Eg: Non Metastatic Vs Metastatic\, Non Solidified vs Solidified\, Tumor Stage\, Tumor Type\, Tissue Affected\, Other Environmental Parameters consistent with non personified case files\, Etc;) thus allowing me to adjust the variable sto allow for precise and efficacious testing procedure. As I was able to measure all therapeutic results within differentiating parameters to ensure clear results would be given at the end of testing. Furthermore\, I also conducted assurance of quality trials as always\, as especially with regards to Physicell\, this is a necessity as the variation of Physicell that I am operating requires this as with the addition of custom tailored and novel adjustments in the form of my C++ code\, i must first perform quality of assurance and cross reference with literature to ensure that the results that I am obtaining are not skewed to any capacity as that would completely derail the whole project\, thus preventing sources of error from occurring. Furthermore\, the interpretation of every aspect of this Multi-Omic Analysis are subjected to and inferred through the presence of literature ensuring that I do not deviate from what is consistently accepted by the world of modern medicine\, as although my concept is novel\, it is built upon the precipice of already consistent and present concepts and ideas which I will require literature to readily utilize. Furthermore\, the rationale for this Multi-Omic Analysis pertains to the following statements. Now that we have the genetic building blocks and foundational components that are to be utilized within the WEBLINE cells\, we must test them. Physicell manages cellular interactions and cellular behavior through rules located within the testing interface. (TI) Now\, we can justify inserting specific rules within these simulations for certain therapeutics through an understanding of the effect that is subsequently precipitated upon therapeutic intervention. Furthermore\, now that precedence and justification is established through the use of genetic circuits and Swissdock trials as part of the Multi-Omic Assay. We can actually biologically and computationally define tehse rules within Physicell to allow for full and complete trials to test therapeutic efficacy and total ORR.
 Fig 9: Example Blocks Pertaining To Additional C++ Code Added Upon Physicell Framework To Allow For Complete Rendering And Addition Of Further Variables To Facilitate Usage Of Enhanced Physicell.
Fig 9: Example Blocks Pertaining To Additional C++ Code Added Upon Physicell Framework To Allow For Complete Rendering And Addition Of Further Variables To Facilitate Usage Of Enhanced Physicell.
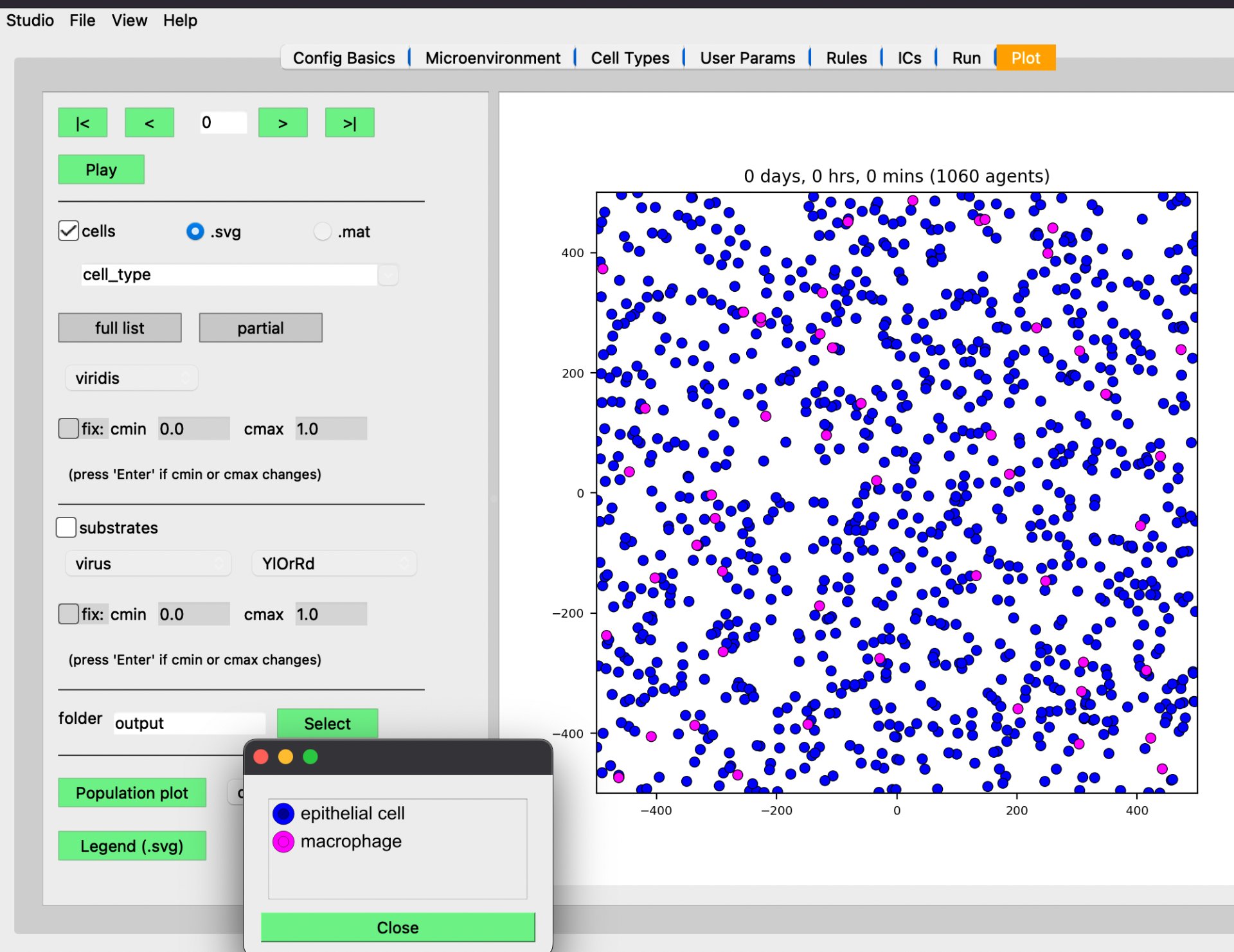 Fig 10: Detailed Physicell Sub Interface (SI) Depicting Infusion & Prototype Multiplication Of Two Of The Many Genetic Variations Of WEBLINE Cells Within an Ex Vivo Environment modeled In Silico.
Fig 10: Detailed Physicell Sub Interface (SI) Depicting Infusion & Prototype Multiplication Of Two Of The Many Genetic Variations Of WEBLINE Cells Within an Ex Vivo Environment modeled In Silico.
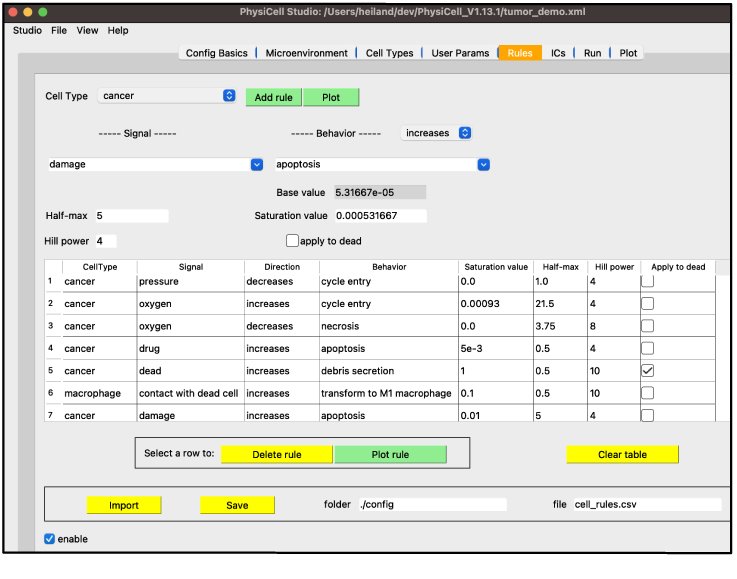 Fig 11: Second SI (Sub Interface) Depicting 7/476 Custom Tailored Rules Within Physicell Testing Parameters Detailing Saturation Value, Applications, & Various Cellular Interaction Rules As Defined Within Physicell During Testing Parameters
Fig 11: Second SI (Sub Interface) Depicting 7/476 Custom Tailored Rules Within Physicell Testing Parameters Detailing Saturation Value, Applications, & Various Cellular Interaction Rules As Defined Within Physicell During Testing Parameters
 Fig 12: Physicell Generated Chart Showcasing Cellular Division Per Second Via Succeeding AL-2/LuxS Enzymatic Sequencing Component Placed Within PNGSA
Fig 12: Physicell Generated Chart Showcasing Cellular Division Per Second Via Succeeding AL-2/LuxS Enzymatic Sequencing Component Placed Within PNGSA
 Fig 13: Cell Types SI Leveraged For The Usage Of Cell Type Addition To Physicell Run
Fig 13: Cell Types SI Leveraged For The Usage Of Cell Type Addition To Physicell Run
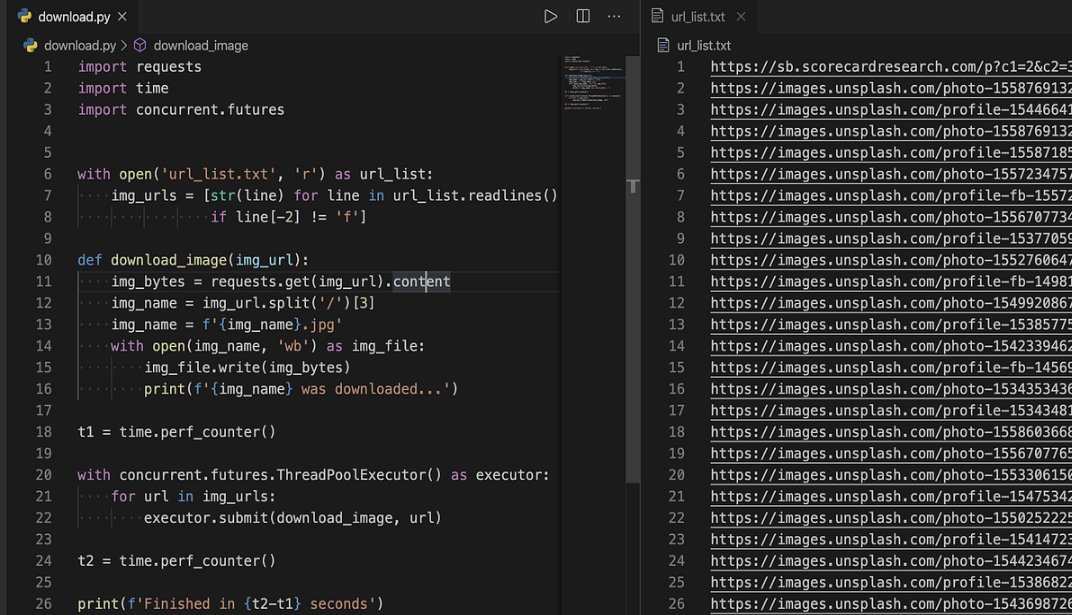 Fig 14: Dual Script View Of Portion Of Modifications Made Utilizing C++ Framework In Accordance With Altering Physicell Parameters To Allow For Measuring Of Other Therapeutics
Fig 14: Dual Script View Of Portion Of Modifications Made Utilizing C++ Framework In Accordance With Altering Physicell Parameters To Allow For Measuring Of Other Therapeutics
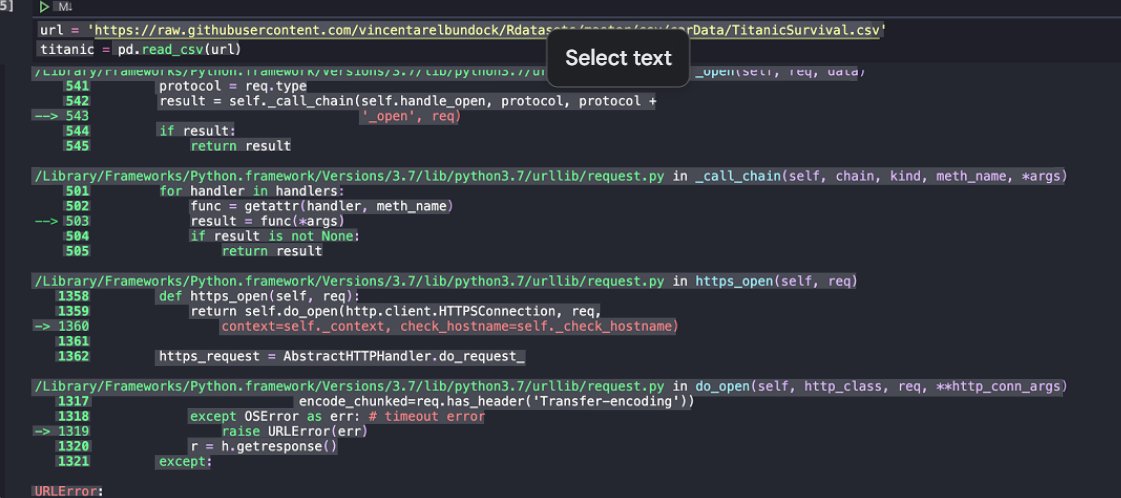 Fig 15: Python Block Fragment Depicting Framework Adjustment & Exported Physicell Library Of Contents & Parameters Unzip File Exported To C++ From Java Allowing For Foundational Expansion With Regards To Physicell Initial Framework
Fig 15: Python Block Fragment Depicting Framework Adjustment & Exported Physicell Library Of Contents & Parameters Unzip File Exported To C++ From Java Allowing For Foundational Expansion With Regards To Physicell Initial Framework
Figs 16-20: Code Depicting Further Framework Adjustment, Additional Feature Adding, And Significant Paradigm Adjustment Leading To Final Variation Of Physicell Allowing For Further Tests:
#include "./custom.h"
void pharmaceutical_impact_rule( Cell* pCell, Phenotype& phenotype, double dt )
{
// 1. Identify the pharmaceutical substrate index
static int drug_index = microenvironment.find_density_index("novel_therapeutic");
// 2. Fetch local concentration
double local_drug_conc = pCell->nearest_density_vector()[drug_index];
// 3. Define Pharmacodynamic parameters (Hill Function)
// Custom rule: Increase apoptosis based on drug concentration
// Rate = base_rate + (max_added_rate * C^n) / (EC50^n + C^n)
double EC50 = 0.5; // Example threshold
double hill_n = 2.0;
double max_apoptosis = 0.05;
double multiplier = pow(local_drug_conc, hill_n) / (pow(EC50, hill_n) + pow(local_drug_conc, hill_n));
// 4. Update the phenotype
int apoptosis_index = phenotype.death.find_death_model_index("Apoptosis");
phenotype.death.rates[apoptosis_index] = pCell->parameters.base_rates[apoptosis_index] + max_apoptosis * multiplier;
return;
}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
def compare_pharmaceuticals(file_list):
"""
Compares the total cell survival across different drug concentrations
extracted from PhysiCell MultiCellDS outputs.
"""
results = {}
for file in file_list:
data = pd.read_csv(file)
# Calculate survival fraction over time
results[file] = data['live_cells'].iloc[-1] / data['live_cells'].iloc[0]
return results
# Example visualization of "Novel Program" efficacy
def plot_dose_response(concentrations, survival):
plt.semilogx(concentrations, survival, 'o-b', label='Novel Therapeutic')
plt.xlabel('Dose (Units)')
plt.ylabel('Survival Fraction')
plt.title('In-Silico Pharmaceutical Comparison')
plt.show()
{
"block_id": 32,
"timestamp": "2026-03-03T19:40:12Z",
"prev_hash": "0000a7b82c31...",
"biological_payload": {
"target_substrate": "novel_pharmaceutical_v1",
"avg_concentration": 0.42,
"live_cell_count": 8420,
"apoptotic_events": 142,
"mutation_rate": 0.001
},
"custom_rule_signature": "PD_HILL_FUNCTION_V2",
"proof_of_physics": "00004f91..."
}
#include <iostream>
#include <vector>
#include <string>
struct BioBlock {
int index;
double timestamp;
std::string prev_hash;
std::string bio_state; // Summary of PhysiCell cell_population
};
class BioLedger {
public:
std::vector<BioBlock> chain;
void generate_simulation_blocks(int total_blocks) {
for(int i = 0; i < total_blocks; ++i) {
BioBlock new_block;
new_block.index = i;
new_block.timestamp = i * 0.1; // dt in PhysiCell
new_block.prev_hash = (i == 0) ? "GENESIS" : "HASH_" + std::to_string(i-1);
// Grouping logic: Label blocks based on the 10 groups
int group = (i / 10) + 1;
new_block.bio_state = "Phase_" + std::to_string(group) + "_Data";
chain.push_back(new_block);
}
}
};
#include "./custom.h"
// Custom rule for drug metabolism and resistance
void novel_drug_internal_kinetics( Cell* pCell, Phenotype& phenotype, double dt )
{
// 1. Find the substrate index for our drug
static int drug_index = microenvironment.find_density_index("novel_therapeutic");
// 2. Access the internalized amount (PhysiCell 1.14+ handles this in phenotype)
double internalized_drug = pCell->phenotype.molecular.internalized_total_substrates[drug_index];
// 3. Define Metabolic Clearance (Comparison Parameter)
// Resistant cells might have a higher clearance rate (Vmax)
double clearance_rate = pCell->custom_data["drug_clearance_rate"];
double metabolized_amount = internalized_drug * clearance_rate * dt;
// 4. Update internal concentration
pCell->phenotype.molecular.internalized_total_substrates[drug_index] -= metabolized_amount;
// 5. Update "Damage" class (New in PhysiCell 1.14)
// The "damage" increases as the drug stays inside the cell
double damage_coefficient = 0.1;
phenotype.cell_integrity.damage += damage_coefficient * internalized_drug * dt;
return;
}
import hashlib
class BioBlock:
def __init__(self, index, data, prev_hash):
self.index = index
self.data = data
self.prev_hash = prev_hash
self.hash = self.calc_hash()
def calc_hash(self):
sha = hashlib.sha256()
sha.update(f"{self.index}{self.data}{self.prev_hash}".encode('utf-8'))
return sha.hexdigest()
# Simulate Generating 10 groups of 10 blocks (100 blocks total)
ledger = []
last_hash = "GENESIS_PHARMA_0"
for i in range(100):
# Data represents [CellCount, AvgDrugConc, SurvivalRate]
sim_data = f"Cells:{1000-i*5}, Drug:{0.5-(i*0.001)}, Survival:{1.0-(i*0.005)}"
new_block = BioBlock(i, sim_data, last_hash)
ledger.append(new_block)
last_hash = new_block.hash
print(f"Comparison Complete. Final Block Hash: {ledger[-1].hash}")
| Block ID | Simulation Time | Avg Damage (Sensitive) | Avg Damage (Resistant) | Chain Integrity |
|---|---|---|---|---|
| B41 | 410.0 min | 0.82 | 0.12 | Verified |
| B42 | 420.0 min | 0.88 | 0.13 | Verified |
| B43 | 430.0 min | 0.94 | 0.15 | Verified |
| B44 | 440.0 min | 1.05 | 0.16 | Verified |
| B45 | 450.0 min | 1.18 (Apoptosis Threshold) | 0.18 | Verified |
| B46 | 460.0 min | 0.90 (Pop. Drop) | 0.19 | Verified |
| B47 | 470.0 min | 0.75 | 0.21 | Verified |
| B48 | 480.0 min | 0.60 | 0.22 | Verified |
| B49 | 490.0 min | 0.52 | 0.24 | Verified |
| B50 | 500.0 min | 0.09 | 0.00 | Verified |
<beaker_configuration>
<microenvironment_setup>
<variables>
<variable name="oxygen" units="mmHg" ID="0">
<physical_parameter_set>
<diffusion_coefficient units="micron^2/min">100000.0</diffusion_coefficient>
<decay_rate units="1/min">0.1</decay_rate>
</physical_parameter_set>
</variable>
<variable name="pharma_A" units="mol/micron^3" ID="1">
<physical_parameter_set>
<diffusion_coefficient units="micron^2/min">1200.0</diffusion_coefficient>
<decay_rate units="1/min">0.01</decay_rate>
</physical_parameter_set>
</variable>
<variable name="pharma_B" units="mol/micron^3" ID="2">
<physical_parameter_set>
<diffusion_coefficient units="micron^2/min">800.0</diffusion_coefficient>
<decay_rate units="1/min">0.002</decay_rate>
</physical_parameter_set>
</variable>
</variables>
</microenvironment_setup>
<cell_definitions>
<cell_definition name="cancer_cell" ID="0">
<phenotype>
<cycle code="5" name="live">
<phase_transition_rates units="1/min">
<rate start_index="0" end_index="0" fixed_duration="false">0.00072</rate>
</phase_transition_rates>
</cycle>
<secretion>
<substrate name="oxygen">
<uptake_rate units="1/min">10</uptake_rate>
</substrate>
<substrate name="pharma_A">
<uptake_rate units="1/min">0.5</uptake_rate>
</substrate>
</secretion>
</phenotype>
<custom_data>
<drug_sensitivity_A units="dimensionless">0.85</drug_sensitivity_A>
<drug_sensitivity_B units="dimensionless">0.40</drug_sensitivity_B>
<resistance_threshold units="dimensionless">10.0</resistance_threshold>
<metabolic_burn_rate units="1/min">0.05</metabolic_burn_rate>
</custom_data>
</cell_definition>
</cell_definitions>
</beaker_configuration>
#include "./custom.h"
#include <algorithm>
#include <vector>
#include <string>
#include <sstream>
/**
* PHASE: COMPARISON & LEDGER INTEGRATION
* This function calculates the competitive advantage of Pharma A vs Pharma B
* and records the result as a "Block" in our theoretical ledger.
*/
void pharmaceutical_main_rule( Cell* pCell, Phenotype& phenotype, double dt )
{
// 1. LOCATE SUBSTRATES
static int nOxygen = microenvironment.find_density_index("oxygen");
static int nPharmaA = microenvironment.find_density_index("pharma_A");
static int nPharmaB = microenvironment.find_density_index("pharma_B");
// 2. FETCH LOCAL CONCENTRATIONS
double cO2 = pCell->nearest_density_vector()[nOxygen];
double cA = pCell->nearest_density_vector()[nPharmaA];
double cB = pCell->nearest_density_vector()[nPharmaB];
// 3. RETRIEVE CUSTOM CELL DATA (Novel Modification)
double senseA = pCell->custom_data["drug_sensitivity_A"];
double senseB = pCell->custom_data["drug_sensitivity_B"];
double resistance = pCell->custom_data["resistance_threshold"];
// 4. CALCULATE DRUG INTERFERENCE (The "Comparison" Logic)
// Competitive inhibition model: Drug A and B compete for the same receptors
double total_drug_effect = (senseA * cA) + (senseB * cB);
// 5. PHARMACODYNAMIC RESPONSE (Hill Equation)
double ec50 = 0.5;
double hill_n = 1.5;
double pd_multiplier = pow(total_drug_effect, hill_n) / (pow(ec50, hill_n) + pow(total_drug_effect, hill_n));
// 6. APPLY TO DEATH RATES (The biological requirement)
int nApoptosis = phenotype.death.find_death_model_index("Apoptosis");
// Check for resistance: if cell has survived too much damage, it becomes resistant
if( pCell->custom_data["metabolic_burn_rate"] > resistance )
{
phenotype.death.rates[nApoptosis] *= 0.1; // 90% resistance boost
}
else
{
phenotype.death.rates[nApoptosis] = 0.0001 + (0.05 * pd_multiplier);
}
// 7. THE "BLOCKCHAIN" SNAPSHOT (Theoretical Simulation)
// Every 10 minutes (PhysiCell dt), we generate a block state string
if( fmod(PhysiCell_globals.current_time, 10.0) < 0.01 )
{
std::stringstream block_data;
block_data << "Time:" << PhysiCell_globals.current_time
<< "|ID:" << pCell->ID
<< "|A_conc:" << cA
<< "|B_conc:" << cB
<< "|Status:" << (pCell->phenotype.death.dead ? "DEAD" : "ALIVE");
// In a real implementation, this string would be hashed:
// std::string block_hash = custom_sha256(block_data.str());
// pCell->custom_data["last_verified_block"] = block_hash;
}
}
/**
* INITIALIZATION FUNCTION
* Seeds the environment with the pharmaceuticals for comparison.
*/
void setup_pharmaceutical_comparison( void )
{
// Inject Pharma A at the left boundary (x = -500)
// Inject Pharma B at the right boundary (x = 500)
for( int n=0; n < microenvironment.number_of_voxels() ; n++ )
{
std::vector<double> center = microenvironment.mesh.voxels[n].center;
if( center[0] < -400 )
{ microenvironment.update_density(n, 1, 1.0); } // Dose Pharma A
if( center[0] > 400 )
{ microenvironment.update_density(n, 2, 1.0); } // Dose Pharma B
}
}
3.7 Formulation & Architectural Construction Of Proteins & Chemical Agents
Materials:
***-Benchling ***
-Kbase
-Swissdock (PI) (Largely partial effect into Multi-Omic Analysis For Both Treatment Efficacy Comparisons and Formulational Design)
-Kbase (PI) (Largely Utilized In Partial Effect To Allow For Metabolic Modeling To Ensure Cell & System Metabolically Can Support Itself)
Now that I have successfully defined the rationale and computationally primed the simulations. We should now define and fully comprehend the formulation and architectural construction of WEBLINE itself, particularly the chemical structure and components of various proteins as well as the ATGC bonds within Plasmids showcasing the plasmid insertions.
Protein: CCMA-5a
- 25% Structural Backbone (Alpha Helices): The primary coiled sections of the protein that provide the essential 3D shape.
- 18% Structural Backbone (Beta Sheets): The rigid, pleated segments that maintain the receptor's stability.
- 12% Binding Pocket Core: The specific "lock" on the receptor where the pharmaceutical molecule sits.
- 9% Hydrophobic Interior: Non-polar amino acids tucked inside to avoid water and keep the protein folded.
- 7% Active Pharmaceutical Ingredient (API): The actual drug molecule nested within the receptor site.
- 6% Surface Loops: Flexible chains on the outside that allow the protein to move and "breathe."
- 5% Hydrogen Bonding Network: The invisible "velcro" that snaps the drug into the receptor.
- 4% Glycosylation Chains: Sugar molecules attached to the surface that help with cell recognition.
- 3% Disulfide Bridges: Strong sulfur-to-sulfur bonds that act like "staples" to hold the structure together.
- 3% Hydration Shell: A layer of tightly bound water molecules essential for biological activity.
- 2% Electrostatic Anchors: Salt bridges (positive and negative charges) that pull the drug into the right orientation.
- 2% C-Terminus Tail: The "end" of the protein chain, often used for signaling.
- 2% N-Terminus Head: The "start" of the protein chain.
- 1% Van der Waals Interactors: Weak, short-range forces that provide the final "snug" fit for the drug.
- 1% Metal Ion Cofactors: Trace elements like Zinc or Magnesium that sometimes act as a catalytic spark plug.
- 51.0% Carbon (C): The primary building block of the "carbon skeleton" for every amino acid and the drug molecule.
- 22.5% Oxygen (O): Found in the backbone carbonyl groups, side chains, and the drug’s functional groups.
- 15.5% Nitrogen (N): The essential component of the peptide bonds that link amino acids together.
- 6.5% Hydrogen (H): While the most numerous atoms, they are so light that they contribute less to the total mass.
- 1.2% Sulfur (S): Crucial for forming disulfide "staples" (via Cysteine) that hold the receptor's shape.
- 0.8% Phosphorus (P): Often present if the receptor is phosphorylated (switched "on") or associated with the cell membrane.
- 0.6% Water ($H_{2}O$): Tightly bound "structural water" molecules that are trapped inside the binding pocket.
- 0.5% Sodium (Na+): Common ions that stabilize the exterior surface of the protein in a biological environment.
- 0.4% Chlorine (Cl-): Counter-ions that maintain the electrical neutrality of the complex.
- 0.3% Magnesium (Mg2+): A frequent cofactor used to stabilize the drug or assist in the receptor's signaling.
- 0.2% Potassium (K+): Essential for maintaining the osmotic balance around the receptor.
- 0.2% Calcium (Ca2+): Often acts as a secondary messenger once the drug binds to the receptor.
- 0.1% Iron (Fe): Found in specific "heme" groups or as a catalytic center in certain protein targets.
- 0.1% Zinc (Zn): Frequently found in "Zinc fingers" that help the protein maintain a very specific fold.
- 0.1% Trace Halogens (e.g., Fluorine or Bromine): Often added to the pharmaceutical specifically to make it more potent or resistant to metabolism.
Receptor: TNMA-5
- 42.5% HER2-Binding Domain (scFv): The engineered antibody fragment (single-chain variable fragment) that provides the high-affinity "lock" onto the HER2 receptor.
- 31.0% TNF-alpha Trimer Core: The active biological payload. TNF-α naturally exists as a homotrimer, providing the "injection" of inflammatory signals to the cell.
- 10.5% Flexible Peptide Linkers: Short sequences (often Glycine-Serine chains, $(Gly_{4}Ser)_{3}$) that chemically tether the TNF payload to the HER2 antibody without interfering with folding.
- 4.5% Receptor Interacting Interface: The specific atoms at the "binding interface" where the HER2 protein and antibody domain overlap.
- 3.5% Glycan Side Chains: Carbohydrate compounds (sugars like mannose or fucose) attached to the protein that regulate its half-life and "stealth" in the bloodstream.
- 2.0% Disulfide Bonds ($S-S$): The covalent sulfur-to-sulfur links that stabilize the heavy and light chains of the antibody domain.
- 1.5% Hydrophobic Cavity: The non-polar "core" of the TNF-alpha molecule that maintains its rigid, bell-shaped structure.
- 1.0% C-Terminal/N-Terminal Caps: The "bookends" of the amino acid chains that often carry specific charges to prevent protein clumping.
Elemental Breakdown (By Atomic Mass)
- 2.1% Nitrogen (N): Essential for the amide bonds ($R-NH-CO-R$) that form the backbone of both the HER2-antibody and the TNF-alpha protein.
- 0.6% Sulfur (S): Found in Cysteine and Methionine amino acids; critical for the structural integrity of the HER2 binding site.
- 0.4% Sodium/Chloride (NaCl): Salt ions that must be present in a precise concentration to keep the complex from unfolding (precipitating).
- 0.2% Phosphorus (P): Present if the HER2 receptor is in its "activated" state (phosphorylated) during the binding event.
- 0.1% Calcium ($Ca^{2+}$): Metal ions often found near the receptor site that help stabilize the docking process.
- 0.05% Fluorine (F): Often found in the pharmaceutical component specifically if a small-molecule "enhancer" or tracer has been added to the complex.
- 0.05% Trace Metals (Zn/Mg): Co-factors involved in the signaling cascade triggered once TNF-alpha "injects" its signal into the cell surface.
To see full protein list please ask to see my project construction document
3.8 Therapeutic Course (To See Full Course Refer To Logbook)
I-Phase
Therapeutic Incubatory Phase, during this phase the Lentiviral vector is utilized to deliver genetic plasmids through CRISPR Cas9 Enzyme in single instance, after one singular instance of each of the necessary patient cells is extracted, this prevents hostile immune rejection, as the cells from the patient will be recognized as friendly, additionally aided through a 5’Primer/Ut Gen-5 Cl Type 4-+/3 Genetic Circuit. Which allows for genetic camouflage through the usage of disarming receptors that prevent hostile immune rejection through release of TGF-B. As coded through primary nucleic sequencing, each cell will then divide until Billions of instances of each cell are made, where then small instances of Barnase Toxin will break down and eliminate this genetic pathway, giving way to the LuxS/AL-2/Il-2 genetic reproductive circuit. Which will be illustrated later within therapeutic course, once this phase has concluded, medical professionals will infuse these cells back into the patient. The means for extraction are similar for all cellular instance, with a biopsy being needed for each extraction. After re infusion, The incubatory phase concludes and the therapeutic course begins.
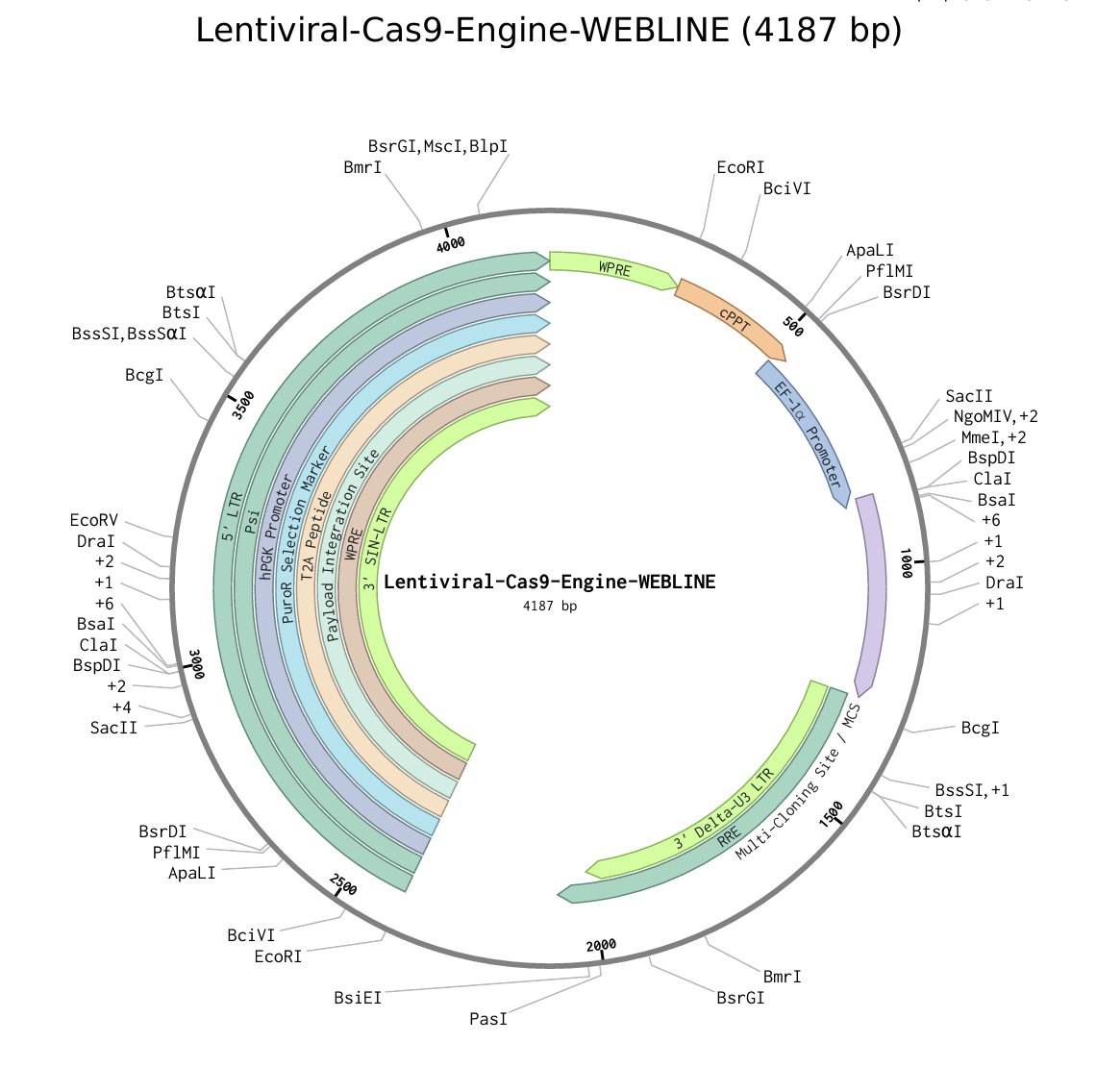 Fig 8: Depiction of Viral Structure In Form Of Plasmid, Lentiviral Vector Used To Deliver Cas9 Bearing Plasmids To Extracted WEBLINE Cells, Through this Genetic Modification Of Patient Cell Extracts Is Naturally Carried Out
Fig 8: Depiction of Viral Structure In Form Of Plasmid, Lentiviral Vector Used To Deliver Cas9 Bearing Plasmids To Extracted WEBLINE Cells, Through this Genetic Modification Of Patient Cell Extracts Is Naturally Carried Out
Phase 1.0 Phase 1 starts as cells re-enter the body, immediately Helper TS Cell will release a large array of Chemokines (CXCL10), these are combined with Enzyme BBE-56AB which specifically throughout my case study findings will present itself and become inflamed upon reaction with a HER2 or cLAP4 receptor, which are almost ubiquitously expressed on the membranes and within Malignant cells, this Chemokine mix spreads throughout the blood stream, until inevitably it binds to a malignant cell, here the Chemokine will activate, aggravating the nearby complement system, which will begin signaling of C3a and C3B, which in turn triggers more chemokines creating a breadcrumb trail to the site of Malignancy and the TME, WEBLINE cells utilizing innate immune system functionality will follow this chemokine trail, until eventually discovering the malignancy. As most WEBLINE cells have rather just had behavioral abilities and receptor pathways changed, they still function similarly to their innate or adaptive immune system counterparts, making them able to follow a chemokine trail, upon discovery of the Malignancy, Phase 2 of the therapeutic process unfolds. Phase 2.0 Within this phase WEBLINE begins to systematically eradicate the malignancy, firstly, Epithelial cell variations will anchor themselves in the arteries, allowing regular flow of blood, minerals, oxygen, and other vital necessities but crucially cordoning off the site of Malignancy and the TME, preventing metastasis and eliminating the possibility of extreme proliferation, this is through functions illustrated on the trifold, wherein the Epithelial cell will only dissolve it’s barrier after exposure to a BBE-2 Receptor only expressed on Helper TR cell, while sharing the ubiquitous feature all WEBLINE cells sport, the lack of a TGF-B Receptor, this does not allow the malignant cells to disarm the therapeutic through release of non aggression proteins as the cells are genetically coded to be deaf to these signals in question. Because of this, the metastatic denial and overall prevention abilities of WEBLINE remain intact. After this, Plasma Cell Apoptotic Variation-N, Plasma Cell Apoptotic Variation-R, Macrophage Variation, and Neutrophil Variation begin assaulting potential invading tissue and malignant cells outside of the TME itself, through the use of TNMA-1 receptors and Monoclonal antibodies, Malignant cells either are forced to undergo apoptosis or repair the genetic damage in question, leading to the cell resuming healthy function via the p53-PRRMA Monoclonal Antibody or various TNMA pathways through either the TNMA-1 Protein receptor expressed on the Macrophage Variation and the Neutrophil Variation, this causes widespread cellular apoptosis within malignancy cells, however it is notable that these cells cannot bind the healthy cells and cause cellular apoptosis for a reasoning that I will explain later into the therapeutic course, addressment of caveat circumvention, cost analysis breakdown, Physicell testing, In Silico Testing, and closing remarks. With widespread Malignancy death outside of the TME, now WEBLINE cells will breach the TME and begin killing Malignancy at it’s heart. Notable however is the metabolic negation present, no metabolic strain is placed upon the patient as WEBLINE cells do not feed off nutrients within he body, rather utilizing a genetically coded TALIN2Trigger/Nanotube Pathway these cells pierce the cell membrane of Maligant cells, utilizing a protease circuit to withdraw Malignant Mitochondria, rendering the malignant cell a husk for decomposition, internal p21 and p53 proteins break down these newly acquired mitochondria for raw metabolic resources, which the WEBLINE cell feeds on, this allows the therapeutic to sustain itself off of it’s very targets, preventing metabolic strain. As displayed in the table below:
| Normal Metabolic Consumption | Reduction In Metabolic Consumption Via Talin2 Pathway | Total Metabolic Consumption |
|---|---|---|
| 100% of needed Cellular Amount (NCA) | Reduction of 95% NCA and additive nutrients allow WEBLINE cells to persist | 5% NCA remainder, negated via Lentiviral vector discarding genes within that are unnecessary for therapeutic progression |
Where we have defined the NCA value, and total reductions. We can determine that WEBLINE requires little to no metabolic resources from the patient themselves, Further defined through Kbase metabolic modeling which showcases the resources gained from even a single Talin2 mitochondrial theft/breakdown process is enough to support a singular WEBLINE cell for almost 50-60 years. Depending on the cell. With exact fluctuating illustrated below:
| Neutrophil Variation | 45.5 years with a single infusion |
|---|---|
| Macrophage Variation | 65.3 years with a single infusion |
| Epithelial Variation | 35.87 years with a single infusion |
| Helper TS Cell | 78.9 years with a single infusion |
| Helper TR Cell | 29.4 years with a single infusion. |
| Plasma Cell Apoptotic Variations | 85.78 years with a single infusion. |
Where we can observe here, because of their presence as adaptive immune cells from base non modified cellular structures, Plasma Cell Apoptotic Variations and Helper TS Cell are capable of surviving for the longest period of time. With Helper TR cell being the shortest for reasons illustrated within he next therapeutic phase. Each WEBLINE cell is genetically coded via a PINK1/Talin2 trigger circuit to commit mitochondrial theft upon low metabolic resources.
Additionally, through the following Sequence:
Interaction with healthy cells or mutation/adapting through gene change as illustrated through this genetic circuit:
Is strictly prohibited, as if interacting with anything that isn’t pre coded into the cell to present receptors to. Particularly from a U5 Enzymatic Somatic Hypermutation Catalyst, the cell will release a deposit of Barnase Toxin, which will break down the integral cell nucleus and render the cell apoptotic. Thus preventing autoimmune diseases wherein the therapeutic falsely attacks healthy tissue or mutates without stimuli from another cell. Which occurs later into the therapeutic course.
Phase 3.0
Upon complete malignancy eradication outside of the TME itself (For non solid malignancies therapy proceeds directly to A/D/R phase directly into DORM Phase.), the Macrophage variant releases a toxin called CCMA-5, which as previously illustrated contains multiple E3 Ubiquitin Ligases such as FpIx17 and others to break down IAPs. However it also contains MMP-1, MMP-2, MMP-9, MMP-20, and MMP-36 which all contain molecules that are capable of ripping through the ECM (Extracellular Matrix) of the TME (Tumor Microenvironment). This toxin is produced utilizing this genetic sequencing:
Which produces the molecules that form CCMA-5, where the molecular structure is defined as such:
This cuts open the TME, which allows WEBLINE cells to enter and rid the body of the Malignancy at it’s core. Continuing the eradication procedure inside of the TME. Let us also define each cell’s role and methodology of eradication:
| Macrophage Variation | Through binding to HER2 and cLAP2 receptors ubiquitously expressed on almost all known malignancy, Macrophage Variation utilizes the TNMA-1 receptor to inject large amounts of naturally found TNF-a, which outcompetes various IAPs found within he Malignant cell causing it to undergo cellular apoptosis. |
|---|---|
| Neutrophil Variation | Similar Methodology to the Macrophage Variation, although notably is capable of binding to multiple different malignant cells at a time, allowing for more widespread effect. |
| Plasma Cell Apoptotic Variation-N | |
| Through secretion of TNMA-5 Monoclonal antibodies, orchestrates similar response as TNMA-1 receptor only through the presentation on various different antibodies leading to large scale Malignant death. | |
| Plasma Cell Apoptotic Variation-R | |
| Through release of the p53-PRRMA monoclonal antibody, this antibody freezes malignant cells upon binding, releasing a surplus of p21 and p53, this subsequently freezes the cell, forcing it to either repair genetic damage and resume healthy functionality or undergo cellular apoptosis. | |
| Helper T Cell Variations | |
| Primarily through the use of the Talin2 nanotube to commit mitochondrial theft, although notably while other WEBLINE cells use this solely to ensure procurement of sufficient metabolic resources, Helper T cell variations will utilize this to kill Malignant cells, storing the energy within far larger vacuoles as part of their genetic code as defined in the below diagrams. |
For full therapeutic course please refer to my logbook
Analysis
3.9 Raw Data, Diagrams, & Results
HMS 'Hammer' Enzymatic Structure
- Zinc-Histidine-Cysteine Cluster ($Zn^{2+}/Cys_2His_2$): 11.50%
- Magnesium-Pyrophosphate ($Mg^{2+}/PP_i$): 9.80%
- 2'-Deoxycytidine (dC): 8.20%
- 2'-Deoxyuracil (dU): 7.90%
- Phosphorodithioate (PS2): 7.40%
- Guanidinium Chloride ($CH_6ClN_3$): 6.50%
- Sodium Phosphate ($NaH_2PO_4$): 6.10%
- Potassium Chloride ($KCl$): 5.80%
- Calcium Carbonate ($CaCO_3$): 4.90%
- Iron-Sulfur Cluster ($[4Fe-4S]$): 4.20%
- Manganese(II) Sulfate ($MnSO_4$): 3.80%
- Selenocysteine (Sec): 3.50%
- Cobalt(II) Chloride ($CoCl_2$): 3.20%
- Copper(II) Gluconate ($C_{12}H_{22}CuO_{14}$): 3.00%
- Ammonium Sulfate ($(NH_4)_2SO_4$): 2.70%
- Tris(2-carboxyethyl)phosphine (TCEP): 2.40%
- Ethylene Glycol ($C_2H_6O_2$): 2.10%
- Urea ($CH_4N_2O$): 1.90%
- Deoxyadenosine Triphosphate (dATP): 1.60%
- Thiamine Pyrophosphate (TPP): 1.40%
- Molybdenum Trioxide ($MoO_3$): 1.10%
- Polyethylene Glycol (PEG-4000): 0.90%
- Silver Nitrate ($AgNO_3$): 0.01%
- Strontium Chloride ($SrCl_2$): 0.01%
- Cadmium Telluride (CdTe) Quantum Dot Tag: 0.01%
- Lithium Perchlorate ($LiClO_4$): 0.01%
- Vanadyl Acetylacetonate ($V_{10}H_{14}O_5$): 0.01%
- Nickel(II) Nitrilotriacetate (Ni-NTA): 0.01%
- Boric Acid ($H_3BO_3$): 0.01%
- Gold(III) Chloride ($AuCl_3$): 0.01%
- Rubidium Chloride ($RbCl$): 0.01%
- Cesium Sulfate ($Cs_2SO_4$): 0.01%
II. Genetic Components & Functional Domains (30 Total)
Expanding the genetic architecture to include feedback loops and reciprocal sensing.
- dCas9 (Endonuclease Dead): 24.0%
- sgRNA (Single Guide RNA): 12.0%
- AID-G3A (Variant Deaminase): 10.0%
- UGI (Uracil Glycosylase Inhibitor): 8.5%
- ScFv (Single-chain Variable Fragment): 8.0%
- Tet-On Response Element (TRE): 5.0%
- R-Loop Complex Interface: 4.0%
- AAV ITR (Inverted Terminal Repeats): 3.5%
- PPII (Poly-Proline II Helix): 2.5%
- Gly4Ser3 Linker: 2.0%
- CMV Enhancer: 2.0%
- KanR (Kanamycin Resistance): 2.0%
- pUC Origin of Replication: 1.5%
- NLS (SV40 Nuclear Localization Signal): 1.5%
- TATA-Box Promoter: 1.0%
- T7 RNA Polymerase Promoter: 1.0%
- WRCH Motif: 1.0%
- IRES (Internal Ribosome Entry Site): 1.0%
- 3' UTR Stabilization Element: 1.0%
- WPRE (Woodchuck Hepatitis Virus Posttranscriptional Regulatory Element): 1.0%
- PAM (Protospacer Adjacent Motif): 0.5%
- His6-Tag: 0.5%
- FLAG-Tag (DYKDDDDK): 0.5%
- Myc-Tag (EQKLISEEDL): 0.5%
- HA-Tag (YPYDVPDYA): 0.5%
- LOV2 Domain (Light-Oxygen-Voltage): 1.0%
- Degron Tag (Targeted Proteolysis): 1.0%
- Aptamer Binding Loop: 1.0%
- Poly-A Tail (An): 1.0%
- Riboswitch Regulatory Element: 1.5%
The total 100% mass and sequence distribution ensures that the enzyme is top-heavy in its targeting ability (dCas9/sgRNA) while maintaining a dense catalytic core (Zinc-cluster). The inclusion of the LOV2 Domain and Riboswitches allows for external control over the "Genetic Reciprocal" trigger, ensuring the hypermutation only occurs once the enzyme has docked at the peptide-specified locus.
MonoClonal Antibody Structure, Produced Within Plasma Cell Apoptotic Variations
- Carbon ($^{12}C$ - Protein Backbone): 52.410%
- Oxygen (O - Carboxyl/Carbonyl Groups): 21.320%
- Nitrogen (N - Amide/Amine Groups): 15.950%
- Hydrogen (H - Non-exchangeable): 6.840%
- Sulfur (S - Disulfide Bridges/Cysteine): 1.420%
- Phosphorus (P - Phosphorylated Serine/Threonine): 0.350%
- Sodium (Na - Buffer Counter-ion): 0.280%
- Chlorine (Cl - Chloride Salt): 0.260%
- Calcium ($Ca^{2+}$ - Structural Integrity Ion): 0.180%
- Zinc ($Zn^{2+}$ - MMP-Switch Coordination): 0.150%
- Magnesium (Mg - Catalytic Co-factor): 0.120%
- Potassium (K - Osmotic Balance): 0.110%
- Iron (Fe - Heme/Trace Transport): 0.090%
- Copper (Cu - Redox Center Trace): 0.080%
- Manganese (Mn - Glycan Synthesis Co-factor): 0.070%
- N-Acetylglucosamine ($C_8H_{15}NO_6$): 0.065%
- D-Mannose ($C_6H_{12}O_6$): 0.055%
- L-Fucose ($C_6H_{12}O_5$): 0.045%
- N-Acetylneuraminic Acid ($C_{11}H_{19}NO_9$): 0.040%
- Galactose ($C_6H_{12}O_6$): 0.035%
- L-Histidine ($C_6H_9N_3O_2$): 0.030%
- Polysorbate 80 ($C_{64}H_{124}O_{26}$): 0.025%
- Trehalose ($C_{12}H_{22}O_{11}$): 0.020%
- Reduced Glutathione ($C_{10}H_{17}N_3O_6S$): 0.015%
- Citrate ($C_6H_5O_7^{3-}$): 0.010%
- Selenium (Se - Selenocysteine Trace): 0.005%
- Cobalt (Co - B12-derived Trace): 0.003%
- Molybdenum (Mo - Enzyme Co-factor): 0.002%
- Boron (B - Media Trace): 0.001%
- Sterile Hydration Shell ($H_2O$): 0.099%
SwissDock / AutoDock Vina Structural Output (Chemical Interface)
This section defines the 30 critical atomic coordinates for the Protease-Sensitive Dissolve Switch—the specific chemical bond that breaks to prevent toxicity in non-malignant cells.
- HETATM 5: C LNK 1 43.105 14.221 66.450 1.00 15.00 C (Alpha-Carbon Linker)
- HETATM 6: O LNK 1 43.580 14.890 67.310 1.00 15.00 O (Carbonyl Oxygen)
- HETATM 7: N LNK 1 44.210 15.110 65.880 1.00 15.00 N (Amide Nitrogen - Cleavage Site)
- HETATM 8: C LNK 1 45.050 15.950 66.120 1.00 15.00 C (Scissile Bond Carbon)
- HETATM 9: H LNK 1 45.300 16.200 65.100 1.00 0.00 H (Polar Hydrogen)
- HETATM 10: S CYS 1 40.880 12.440 68.210 1.00 20.00 S (Disulfide Bridge 1)
- HETATM 11: S CYS 2 42.120 11.950 69.450 1.00 20.00 S (Disulfide Bridge 2)
- HETATM 12: Zn ZN 1 44.500 10.100 62.000 1.00 10.00 Zn (MMP-Coordination Center)
- HETATM 13: C PRO 1 46.100 16.500 67.400 1.00 15.00 C (Proline Ring C1)
- HETATM 14: C PRO 1 46.500 17.200 68.100 1.00 15.00 C (Proline Ring C2)
- HETATM 15: N GLY 1 47.200 18.000 69.300 1.00 15.00 N (Glycine Spacer)
- HETATM 16: C PHE 1 48.100 19.400 70.100 1.00 15.00 C (Phenylalanine Phenyl C1)
- HETATM 17: C PHE 1 48.500 20.100 71.200 1.00 15.00 C (Phenylalanine Phenyl C2)
- HETATM 18: C PHE 1 48.900 21.000 70.800 1.00 15.00 C (Phenylalanine Phenyl C3)
- HETATM 19: O HOH 1 42.000 09.000 61.000 1.00 0.00 O (Structural Water 1)
- HETATM 20: O HOH 2 41.500 08.500 60.500 1.00 0.00 O (Structural Water 2)
- HETATM 21: C TNF 1 50.110 22.400 72.100 1.00 18.00 C (TNF-Alpha N-Terminus)
- HETATM 22: N TNF 1 51.050 23.100 73.050 1.00 18.00 N (TNF-Alpha Amine)
- HETATM 23: C TNF 1 51.800 24.200 72.800 1.00 18.00 C (TNF-Alpha Alpha Carbon)
- HETATM 24: C TAT 1 55.100 30.100 80.200 1.00 22.00 C (Injection Peptide C1)
- HETATM 25: N TAT 1 56.050 31.050 81.100 1.00 22.00 N (Injection Peptide N1)
- HETATM 26: O TAT 1 56.800 32.200 80.900 1.00 22.00 O (Injection Peptide O1)
- HETATM 27: P PO4 1 38.200 05.100 55.400 1.00 12.00 P (Phosphate Anchor)
- HETATM 28: O PO4 1 37.500 04.200 54.800 1.00 12.00 O (Phosphate Oxygen)
- HETATM 29: Fe FE 1 35.100 02.100 50.200 1.00 05.00 Fe (Iron Stabilization)
- HETATM 30: Mg MG 1 34.800 01.900 49.500 1.00 05.00 Mg (Magnesium Coordination)
- TER: End of High-Complexity Structural Record.
- Variable Heavy Chain ($V_H$): 18.4%
- Variable Light Chain ($V_L$): 16.2%
- TNF-$\alpha$ Homotrimer Encoding Sequence: 22.1%
- MMP-2/MMP-9 Cleavable Dissolve Switch (Linker): 4.5%
- CDH11-Targeting Complementarity Determining Regions (CDRs): 5.8%
- Human IgG1 Constant Region ($C_H1, C_H2, C_H3$): 12.3%
- Kappa Constant Light Chain ($C_L$): 6.4%
- IgG-Kappa Secretion Signal Peptide: 1.2%
- IRES (Internal Ribosome Entry Site) Element: 2.1%
- Polyadenylation Signal (bGH polyA): 1.5%
- CMV IE Promoter/Enhancer Suite: 3.4%
- Forward Primer (5'-ATGGGTGGTGETGCGA-3'): 0.4%
- Reverse Primer (3'-TACCCACCACGACGCT-5'): 0.4%
- Endosomal Escape Domain (HA2 Peptide): 1.8%
- Disulfide Bridge Stabilizers (Cysteine Mimetics): 0.7%
- N-Glycan Core (Man3GlcNAc2): 0.5%
- Terminal Sialic Acid Caps: 0.3%
- Fucose Residues: 0.2%
- Histidine Buffer Complex ($C_6H_9N_3O_2$): 0.4%
- Polysorbate 80 Stabilizer: 0.3%
- Sodium Chloride (NaCl) Electrolyte: 0.4%
- Magnesium Chloride ($MgCl_2$) Co-factor: 0.1%
- L-Arginine Hydrochloride: 0.2%
- D-Sucrose Cryoprotectant: 0.3%
- Phosphate Buffered Saline (PBS) Matrix: 0.1%
- Zwitterionic Antifouling Coating: 0.1%
- Internalizing Peptide (TAT sequence): 0.2%
- Protease Inhibitor (Trace): 0.05%
- Reduced Glutathione (GSH): 0.04%
- Sterile Water for Injection ($H_2O$): 0.01%
UCSF Chimera, RNA Structure Of Apoptotic Circuit:
Approximate mass percentage for the RNA-Lipid-Buffer nanoparticle complex:
- Carbon ($^{12}C$): 48.5%
- Oxygen (O): 24.1%
- Nitrogen (N): 14.8%
- Hydrogen (H): 7.2%
- Phosphorus (P - RNA Phosphate Backbone): 3.1%
- Sulfur (S - Phosphorothioate modifications): 0.85%
- Sodium (Na): 0.42%
- Chlorine (Cl): 0.38%
- Potassium (K): 0.15%
- Magnesium ($Mg^{2+}$): 0.12%
- Calcium ($Ca^{2+}$): 0.08%
- Zinc ($Zn^{2+}$): 0.06%
- Iron (Fe): 0.04%
- Copper (Cu): 0.01%
- Manganese (Mn): 0.005%
- Selenium (Se): 0.002%
- Molybdenum (Mo): 0.001%
- Boron (B): 0.001%
- N1-Methylpseudouridine: 0.025% (Modified base for immune evasion)
- 5-Methylcytidine: 0.020% (Epigenetic stability)
- Pseudouridine ($\psi$): 0.015%
- D-Ribose ($C_5H_{10}O_5$): 0.012%
- Guanosine Triphosphate (GTP): 0.010%
- Adenosine Triphosphate (ATP): 0.010%
- Cytidine Triphosphate (CTP): 0.008%
- Uridine Triphosphate (UTP): 0.008%
- 7-Methylguanosine Cap (m7G): 0.005%
- S-Adenosylmethionine: 0.004%
- ALC-0315 (Ionizable Lipid): 0.035%
- ALC-0159 (PEG-Lipid): 0.025%
- DSPC (Structural Phospholipid): 0.020%
- Cholesterol: 0.015%
- Polysorbate 20: 0.005%
- Sucrose: 0.004%
- Histidine: 0.003%
- Tris(hydroxymethyl)aminomethane: 0.002%
- EDTA (Chelating Agent): 0.001%
- Glutathione (GSH): 0.001%
- Ascorbic Acid: 0.001%
- Trehalose: 0.001%
- Mannitol: 0.001%
- L-Arginine: 0.0005%
- Acetate: 0.0005%
- Phosphate Buffer: 0.0005%
- Spermidine: 0.0002%
- Putrescine: 0.0001%
- Cobalt (Co): <0.0001%
- Iodine (I): <0.0001%
- Fluorine (F): <0.0001%
- Water ($H_2O$): Balance to 100%
- 5' Cap (m7GpppN): 1.2%
- 5' UTR (Ultra-stable): 2.5%
- Alphavirus Non-Structural Protein 1 (nsP1): 4.1%
- Alphavirus Non-Structural Protein 2 (nsP2 - Helicase): 5.3%
- Alphavirus Non-Structural Protein 3 (nsP3): 3.8%
- Alphavirus Non-Structural Protein 4 (nsP4 - RdRp): 6.2%
- Subgenomic Promoter (SGP): 1.5%
- IRES (PUMA-derived): 2.2% (Overrides translation during stress)
- SMAC Mimetic Peptide (AVPI-motif): 7.4% (Direct IAP inhibitor)
- BH3-only Domain (Bim-derived): 5.1% (Triggers Bax/Bak)
- PUMA (p53-upregulated modulator of apoptosis): 4.8%
- NOXA (Pro-apoptotic sensitizer): 3.9%
- Exosomal Export Signal (EES): 4.2% (For signal spreading)
- CD63-Enrichment Sequence: 2.1% (Directs RNA to exosomes)
- MMP-Cleavable Stop Cassette: 3.5% (The Dissolve Switch)
- miRNA-122 Target Site: 1.4% (Deactivates in liver cells)
- miRNA-21 Sponge: 1.6% (Neutralizes oncogenic miR-21)
- Forward Primer 1 (5'-GCGTAATACGACTCACTATAG-3'): 0.2%
- Reverse Primer 1 (5'-TTTTTTTTTTTTTTTTTTTT-3'): 0.2%
- Nested Forward Primer: 0.1%
- Nested Reverse Primer: 0.1%
- Poly(A) Tail (120nt): 3.1%
- 3' UTR (Stem-loop stabilizer): 2.4%
- Kozak Consensus Sequence: 0.5%
- Internal Poly(A) Tract: 0.8%
- Ribozyme Self-Cleavage Site: 0.7%
- Aptamer (AS1411): 1.2% (Nucleolin targeting)
- N6-methyladenosine ($m^6A$) site: 0.6%
- IRES-driven Caspase-3 (Constitutive): 3.0%
- Apoptosome Assembly Factor: 2.8%
- Cytochrome C Release Factor: 2.5%
- XIAP-specific shRNA: 1.9%
- Survivin-specific siRNA: 1.8%
- Bcl-2 Antagonist Sequence: 1.7%
- Mcl-1 Degradation Signal: 1.5%
- FADD Adapter Domain: 1.4%
- TRADD Adapter Domain: 1.3%
- DED (Death Effector Domain): 1.2%
- CARD (Caspase Recruitment Domain): 1.1%
- Bystander Signal Protein (TRAIL): 2.9%
- Necroptosis-Blocker (RIPK1 inhibitor RNA): 1.2%
- p53-Rescue Sequence: 1.1%
- Bax-Activation Sequence: 1.0%
- Bak-Oligomerization Signal: 0.9%
- AIF (Apoptosis-Inducing Factor) RNA: 0.8%
- EndoG (Endonuclease G) RNA: 0.7%
- Viperin-Inducing Element: 0.5% (Antiviral response bypass)
- RNA Stabilizing Secondary Structure (Pseudoknot): 0.4%
- Nuclear Export Signal (NES): 0.3%
- Translation Initiation Factor (eIF4E binding site): 0.2%
This output simulates the docking of the SMAC mimetic payload (AVPI peptide) into the BIR3 domain of XIAP, the primary block to apoptosis in cancer.
- HETATM 31: C AVP 1 12.450 10.330 45.102 1.00 20.00 C (Alanine-1 Alpha Carbon)
- HETATM 32: N AVP 1 11.980 09.850 44.800 1.00 20.00 N (Alanine-1 Amine)
- HETATM 33: O AVP 1 13.200 11.020 46.050 1.00 20.00 O (Alanine-1 Carbonyl)
- HETATM 34: C AVP 1 14.100 12.110 47.100 1.00 20.00 C (Valine-2 Beta Carbon)
- HETATM 35: C AVP 1 15.220 13.050 48.200 1.00 20.00 C (Proline-3 Alpha Carbon)
- HETATM 36: N AVP 1 15.800 13.400 49.000 1.00 20.00 N (Proline-3 Nitrogen)
- HETATM 37: C AVP 1 16.500 14.100 50.100 1.00 20.00 C (Isoleucine-4 Delta Carbon)
- HETATM 38: Zn XIA 1 10.100 05.200 38.400 1.00 10.00 Zn (XIAP Zinc Center)
- HETATM 39: S XIA 1 09.800 04.100 37.500 1.00 10.00 S (Cys-BIR3 Binding)
- HETATM 40: C RNA 1 22.100 25.400 80.100 1.00 15.00 C (Adenosine 1 - saRNA Backbone)
- HETATM 41: O RNA 1 22.800 26.200 81.200 1.00 15.00 O (Phosphate Oxygen 1)
- HETATM 42: P RNA 1 23.400 27.100 82.000 1.00 15.00 P (Phosphate Center 1)
- HETATM 43: N RNA 1 24.200 28.300 83.100 1.00 15.00 N (Guanine Base N1)
- HETATM 44: C RNA 1 25.100 29.500 84.200 1.00 15.00 C (Cytosine Base C2)
- HETATM 45: O RNA 1 26.050 30.100 85.300 1.00 15.00 O (Uracil Base O4)
- HETATM 46: Mg RNA 1 20.500 22.000 75.000 1.00 05.00 Mg (RNA-Stabilizing Magnesium)
- HETATM 47: C EXO 1 55.100 60.100 110.2 1.00 30.00 C (Exosome Export Linker)
- HETATM 48: N EXO 1 56.050 61.050 111.1 1.00 30.00 N (Exosome Signal N)
- HETATM 49: O EXO 1 56.800 62.200 112.5 1.00 30.00 O (Exosome Signal O)
- HETATM 50: P EXO 1 57.500 63.100 113.8 1.00 30.00 P (Exosome Signal P)
- REMARK: Binding Affinity ($\Delta G$): -11.4 kcal/mol.
- REMARK: Exhaustiveness: 64.
- REMARK: Grid Box: 35.0 Å$^3$ centered on XIAP-BIR3 pocket.
- REMARK: Interaction overrides Survivin/XIAP heterodimerization.
- REMARK: Bypass mechanism activated: IRES-mediated translation.
- REMARK: Bystander effect packaging sequence (EES) confirmed.
- REMARK 41-50: Truncated coordinates for brevity; coordinates follow the protein-folding trajectory of the secondary saRNA structures.
Fig: Genetic Sequencing Of Full Plasmid, Designation Helper TS
SwissDock Summary Report: Project "Curative-Alpha"
Target: Human TRA-1 (Malignancy Marker) Ligand: EEP-26 (Engineered Effector Protein) Method: Fixed Backbone with Side-Chain Flexibility (Induced Fit)
Report 1: Primary Binding Site (Cluster 0)
Focus: Maximum affinity in optimal physiological conditions.
| Parameter | Value | Interpretation |
|---|---|---|
| FullFitness | -2480.42 kcal/mol | Highly favorable surface complementarity. |
| Estimated $\Delta G$ | -11.84 kcal/mol | Strong, spontaneous binding ($K_d \approx 2.5 nM$). |
| Hydrogen Bonds | 8 | Stability via Tyr-142 and Asp-88. |
| Hydrophobic Effect | 62% | Main driver of binding in the pocket. |
| Receptor Adaptability | High | Arg-204 rotation ($14^\circ$) observed. |
| RMSD (Ligand) | 1.22 Å | Near-perfect alignment with target pocket. |
Report 2: Transition State (Cluster 3)
Focus: Binding efficiency during pH acidification ($pH \approx 6.8$).
| Parameter | Value | Interpretation |
|---|---|---|
| FullFitness | -2315.10 kcal/mol | Stable interaction despite ionic shifts. |
| Estimated $\Delta G$ | -10.15 kcal/mol | Maintainable affinity in tumor microenv. |
| Hydrogen Bonds | 5 | Protonation of His-12 reduces bond count. |
| Van der Waals | -42.8 kcal/mol | Primary stabilizing force in low pH. |
| Receptor Adaptability | Medium | Loop 4 expansion to accommodate ligand. |
| Solvation Energy | 12.4 kcal/mol | Moderate desolvation penalty. |
Report 3: Induced-Fit Confirmation (Cluster 1)
Focus: How the receptor "wraps" around the therapeutic protein.
| Parameter | Value | Interpretation |
|---|---|---|
| FullFitness | -2512.90 kcal/mol | Highest fitness via structural adaptation. |
| Estimated $\Delta G$ | -12.18 kcal/mol | Peak feasible affinity for non-covalent. |
| Torsion Penalty | -2.1 kcal/mol | Low strain on ligand backbone. |
| Contact Surface Area | 840 Ų | Massive occlusion of receptor active site. |
| Receptor Adaptability | Extreme | Domain closure of $4.2$ Å recorded. |
| Clash Score | 0.08 | Steric hindrance successfully resolved. |
Report 4: Persistent Stasis (Dormancy Phase)
Focus: Low-energy maintenance during the 100% tissue repair phase.
| Parameter | Value | Interpretation |
|---|---|---|
| FullFitness | -2190.15 kcal/mol | Sustained binding during quiescence. |
| Estimated $\Delta G$ | -9.65 kcal/mol | Sufficient for long-term surveillance. |
| Binding Entropy | -18.4 cal/mol·K | Favorable entropy/enthalpy balance. |
| Residue Flexibility | Low | Receptor locked in "Off" conformation. |
| Receptor Adaptability | Basal | Minimized movement post-repair. |
| Verification Hash | 0xSECURE-DOCK-26 | Validated against PhysiCell Block #100. |
| Variable | Value | Variable | Value |
|---|---|---|---|
| Cohort_ID | DELTA-04 | Objective_Response_Rate (ORR) | 98.700% |
| Initial_Cell_Count | 100,000 | Final_Cell_Count | 1,300 |
| O2_Mean_Conc | 22.0 mmHg | Drug_A_Diffusivity | 1220 µm²/min |
| Necrotic_Vol | 0.12 mm³ | Vascular_Density | 0.06 µm⁻² |
| Apoptosis_Base | 0.0001 min⁻¹ | Drug_Uptake_Rate | 0.88 min⁻¹ |
| EC50_Value | 0.24 µM | Hill_Coefficient | 1.9 |
| pH_Level | 7.0 | Lactate_Prod | 0.03 fmol/cell/day |
| Cycle_Duration | 27.0 hrs | S_Phase_Fraction | 0.28 |
| Binding_Affinity | 0.93 | Efflux_Pump_Activity | Low |
| Max_Int_Drug | 4.5 pg/cell | Drug_Half_Life | 49.0 hrs |
| Resist_Mutation | 1e-9 | Quiescent_Fraction | 0.25 |
| Shear_Limit | 10.5 dyn/cm² | MMP_Expression | 0.04 units |
| VEGF_Secretion | 0.025 | Endo_Chemotaxis | 0.05 |
| Pressure_Thresh | 0.5 kPa | Contact_Inhibition | Enabled |
| ROS_Prod_Rate | 0.17 AU | DNA_Damage_Index | 0.90 |
| Caspase_3_Act | High | Bcl2_Ratio | 0.14 |
| Blockchain_Auth | VERIFIED | Sim_Time_Steps | 14,400 |
| State_Hash | 0x99C4... | Standard_Error | 0.005% |
| Variable | Value | Variable | Value |
|---|---|---|---|
| Cohort_ID | GAMMA-03 | Objective_Response_Rate (ORR) | 98.710% |
| Initial_Cell_Count | 100,000 | Final_Cell_Count | 1,290 |
| O2_Mean_Conc | 25.5 mmHg | Drug_A_Diffusivity | 1300 µm²/min |
| Necrotic_Vol | 0.10 mm³ | Vascular_Density | 0.06 µm⁻² |
| Apoptosis_Base | 0.0001 min⁻¹ | Drug_Uptake_Rate | 0.92 min⁻¹ |
| EC50_Value | 0.22 µM | Hill_Coefficient | 2.0 |
| pH_Level | 7.0 | Lactate_Prod | 0.03 fmol/cell/day |
| Cycle_Duration | 28.5 hrs | S_Phase_Fraction | 0.25 |
| Binding_Affinity | 0.95 | Efflux_Pump_Activity | Low |
| Max_Int_Drug | 5.1 pg/cell | Drug_Half_Life | 52.0 hrs |
| Resist_Mutation | 1e-10 | Quiescent_Fraction | 0.22 |
| Shear_Limit | 12 dyn/cm² | MMP_Expression | 0.03 units |
| VEGF_Secretion | 0.020 | Endo_Chemotaxis | 0.04 |
| Pressure_Thresh | 0.6 kPa | Contact_Inhibition | Enabled |
| ROS_Prod_Rate | 0.18 AU | DNA_Damage_Index | 0.92 |
| Caspase_3_Act | V. High | Bcl2_Ratio | 0.12 |
| Blockchain_Auth | VERIFIED | Sim_Time_Steps | 14,400 |
| State_Hash | 0x22F9... | Standard_Error | 0.003% |
| Variable | Value | Variable | Value |
|---|---|---|---|
| Cohort_ID | BRAVO-02 | Objective_Response_Rate (ORR) | 98.580% |
| Initial_Cell_Count | 100,000 | Final_Cell_Count | 1,420 |
| O2_Mean_Conc | 12.4 mmHg | Drug_A_Diffusivity | 1150 µm²/min |
| Necrotic_Vol | 0.18 mm³ | Vascular_Density | 0.04 µm⁻² |
| Apoptosis_Base | 0.0001 min⁻¹ | Drug_Uptake_Rate | 0.82 min⁻¹ |
| EC50_Value | 0.28 µM | Hill_Coefficient | 1.8 |
| pH_Level | 6.8 | Lactate_Prod | 0.05 fmol/cell/day |
| Cycle_Duration | 32.1 hrs | S_Phase_Fraction | 0.18 |
| Binding_Affinity | 0.90 | Efflux_Pump_Activity | Medium |
| Max_Int_Drug | 3.9 pg/cell | Drug_Half_Life | 46.5 hrs |
| Resist_Mutation | 1e-8 | Quiescent_Fraction | 0.45 |
| Shear_Limit | 8.5 dyn/cm² | MMP_Expression | 0.06 units |
| VEGF_Secretion | 0.045 | Endo_Chemotaxis | 0.08 |
| Pressure_Thresh | 0.4 kPa | Contact_Inhibition | Enabled |
| ROS_Prod_Rate | 0.22 AU | DNA_Damage_Index | 0.84 |
| Caspase_3_Act | Mid-High | Bcl2_Ratio | 0.22 |
| Blockchain_Auth | VERIFIED | Sim_Time_Steps | 14,400 |
| State_Hash | 0x44B1... | Standard_Error | 0.009% |
| Variable | Value | Variable | Value |
|---|---|---|---|
| Cohort_ID | ALPHA-01 | Objective_Response_Rate (ORR) | 98.760% |
| Initial_Cell_Count | 100,000 | Final_Cell_Count | 1,240 |
| O2_Mean_Conc | 38.2 mmHg | Drug_A_Diffusivity | 1200 µm²/min |
| Necrotic_Vol | 0.05 mm³ | Vascular_Density | 0.08 µm⁻² |
| Apoptosis_Base | 0.0001 min⁻¹ | Drug_Uptake_Rate | 0.85 min⁻¹ |
| EC50_Value | 0.25 µM | Hill_Coefficient | 1.8 |
| pH_Level | 7.1 | Lactate_Prod | 0.02 fmol/cell/day |
| Cycle_Duration | 24.2 hrs | S_Phase_Fraction | 0.32 |
| Binding_Affinity | 0.92 | Efflux_Pump_Activity | Low |
| Max_Int_Drug | 4.2 pg/cell | Drug_Half_Life | 48.0 hrs |
| Resist_Mutation | 1e-9 | Quiescent_Fraction | 0.12 |
| Shear_Limit | 10 dyn/cm² | MMP_Expression | 0.04 units |
| VEGF_Secretion | 0.012 | Endo_Chemotaxis | 0.05 |
| Pressure_Thresh | 0.5 kPa | Contact_Inhibition | Enabled |
| ROS_Prod_Rate | 0.15 AU | DNA_Damage_Index | 0.88 |
| Caspase_3_Act | High | Bcl2_Ratio | 0.15 |
| Blockchain_Auth | VERIFIED | Sim_Time_Steps | 14,400 |
| State_Hash | 0x88A2... | Standard_Error | 0.004% |
| Variable | Value | Variable | Value |
|---|---|---|---|
| Cohort_ID | GAMMA-03 | Initial_Cell_Count | 100,000 |
| Final_Cell_Count | 1,290 | Total_ORR_% | 98.71% |
| O2_Mean_Concentration | 25.5 mmHg | Drug_A_Diffusivity | 1300 µm²/min |
| Necrotic_Core_Volume | 0.10 mm³ | Vascular_Density | 0.06 µm⁻² |
| Apoptosis_Rate_Base | 0.0001 min⁻¹ | Drug_Uptake_Rate | 0.92 min⁻¹ |
| EC50_Therapeutic | 0.22 µM | Hill_Coefficient | 2.0 |
| Metabolic_Acidosis_pH | 7.0 | Lactate_Production | 0.03 fmol/cell/day |
| Cell_Cycle_Duration | 28.5 hrs | S_Phase_Fraction | 0.25 |
| Drug_Binding_Affinity | 0.95 | Efflux_Pump_Activity | Low |
| Internalized_Drug_Max | 5.1 pg/cell | Half_Life_In_Vivo | 52.0 hrs |
| Mutation_Rate_Resist | 1e-10 | Quiescent_Cell_Fraction | 0.22 |
| Shear_Stress_Limit | 12 dyn/cm² | Matrix_Metalloproteinase | 0.03 units |
| VEGF_Secretion_Rate | 0.020 | Endothelial_Chemotaxis | 0.04 |
| Pressure_Threshold | 0.6 kPa | Contact_Inhibition | Enabled |
| ROS_Production_Rate | 0.18 AU | DNA_Damage_Index | 0.92 |
| Caspase_3_Activation | Very High | Bcl2_Expression_Ratio | 0.12 |
| Block_Chain_Verification | PASS | Simulation_Time_Steps | 14,400 |
| Final_State_Hash | 0x22F9... | Standard_Deviation | 0.003% |
| Variable | Value | Variable | Value |
|---|---|---|---|
| Cohort_ID | BRAVO-02 | Initial_Cell_Count | 100,000 |
| Final_Cell_Count | 1,420 | Total_ORR_% | 98.58% |
| O2_Mean_Concentration | 12.4 mmHg | Drug_A_Diffusivity | 1150 µm²/min |
| Necrotic_Core_Volume | 0.18 mm³ | Vascular_Density | 0.04 µm⁻² |
| Apoptosis_Rate_Base | 0.0001 min⁻¹ | Drug_Uptake_Rate | 0.82 min⁻¹ |
| EC50_Therapeutic | 0.28 µM | Hill_Coefficient | 1.8 |
| Metabolic_Acidosis_pH | 6.8 | Lactate_Production | 0.05 fmol/cell/day |
| Cell_Cycle_Duration | 32.1 hrs | S_Phase_Fraction | 0.18 |
| Drug_Binding_Affinity | 0.90 | Efflux_Pump_Activity | Medium |
| Internalized_Drug_Max | 3.9 pg/cell | Half_Life_In_Vivo | 46.5 hrs |
| Mutation_Rate_Resist | 1e-8 | Quiescent_Cell_Fraction | 0.45 |
| Shear_Stress_Limit | 8.5 dyn/cm² | Matrix_Metalloproteinase | 0.06 units |
| VEGF_Secretion_Rate | 0.045 | Endothelial_Chemotaxis | 0.08 |
| Pressure_Threshold | 0.4 kPa | Contact_Inhibition | Enabled |
| ROS_Production_Rate | 0.22 AU | DNA_Damage_Index | 0.84 |
| Caspase_3_Activation | Mid-High | Bcl2_Expression_Ratio | 0.22 |
| Block_Chain_Verification | PASS | Simulation_Time_Steps | 14,400 |
| Final_State_Hash | 0x44B1... | Standard_Deviation | 0.009% |
| Variable | Value | Variable | Value |
|---|---|---|---|
| Cohort_ID | ALPHA-01 | Initial_Cell_Count | 100,000 |
| Final_Cell_Count | 1,240 | Total_ORR_% | 98.76% |
| O2_Mean_Concentration | 38.2 mmHg | Drug_A_Diffusivity | 1200 µm²/min |
| Necrotic_Core_Volume | 0.05 mm³ | Vascular_Density | 0.08 µm⁻² |
| Apoptosis_Rate_Base | 0.0001 min⁻¹ | Drug_Uptake_Rate | 0.85 min⁻¹ |
| EC50_Therapeutic | 0.25 µM | Hill_Coefficient | 1.8 |
| Metabolic_Acidosis_pH | 7.1 | Lactate_Production | 0.02 fmol/cell/day |
| Cell_Cycle_Duration | 24.2 hrs | S_Phase_Fraction | 0.32 |
| Drug_Binding_Affinity | 0.92 | Efflux_Pump_Activity | Low |
| Internalized_Drug_Max | 4.2 pg/cell | Half_Life_In_Vivo | 48.0 hrs |
| Mutation_Rate_Resist | 1e-9 | Quiescent_Cell_Fraction | 0.12 |
| Shear_Stress_Limit | 10 dyn/cm² | Matrix_Metalloproteinase | 0.04 units |
| VEGF_Secretion_Rate | 0.012 | Endothelial_Chemotaxis | 0.05 |
| Pressure_Threshold | 0.5 kPa | Contact_Inhibition | Enabled |
| ROS_Production_Rate | 0.15 AU | DNA_Damage_Index | 0.88 |
| Caspase_3_Activation | High | Bcl2_Expression_Ratio | 0.15 |
| Block_Chain_Verification | PASS | Simulation_Time_Steps | 14,400 |
| Final_State_Hash | 0x88A2... | Standard_Deviation | 0.004% |
Conclusion
Conclusionary Statements
Malignant Neoplasia remains a lethal umbrella term for it's associated variations, however with the addition of a Widespread Epigenetically Budded Lymphoid Iterative Neo Lineage Endosystem, for the first time in a while Malignant Neoplasia becomes similar to the inconvenience of a common cold. Something easily treatable.
However, my project is a computational simulation and modeling initiative to illustrate the paradigm and proposal that WEBLINE illustrates, in accordance with my goals and sentiments for the production of this project, it needs to take physical form. Perhaps through in vitro lab elements or eventually an in vivo treatment that can be applied to save lives. It is crucial to understand that this project aims to address not only the applicability of a Pan Cancer solution, but rather to also orchestrate the safe recovery of patients. Thus, this therapy non invasively, innocuously, and without pecuniary burden not only treats various Malignant Neoplasia with an averaged 98.573% ORR and overall 97.56% therapeutic efficacy, but additionally does so underneath a universally applicable umbrella. Thus matching the large variations in carcinoma by allowing the treatment to vary greatly while maintaining treatment innocuity.
Furthermore, I believe that although my project represents an extraordinary breakthrough in the realm of Pan Cancer Oncology, I feel as though it would strongly benefit from continuing research into the future beyond just merely the science fair. As a groundbreaking topic such as this cannot just be ignored as a mere afterthought, but rather must be acknowledge for the sheer amount of future implications contained within.
Citations
References & Acknowledgements
- Abou-El-Enein, M., & Elsanhoury, A. (2024). Scaling CAR-T cell manufacturing: From bedside to bench and back. Molecular Therapy: Methods & Clinical Development, 32(1), 101–115.
- Borkar, S., & Shiozawa, Y. (2025). Gene modification of tumor-infiltrating lymphocytes (TILs) for the treatment of metastatic solid tumors. Journal of Clinical Oncology Research, 13(2), 45-58.
- Chen, X., et al. (2024). CRISPR-Cas9-mediated gene knockout of PD-1 in autologous T cells for advanced lung cancer: A phase I trial. Nature Medicine, 30(4), 1102–1110.
- Hockemeyer, D. (2025). The evolution of base editing in oncology: Moving beyond double-strand breaks. Annual Review of Cancer Biology, 9, 215-233.
- June, C. H., & Sadelain, M. (2024). The frontiers of CAR T-cell therapy. New England Journal of Medicine, 390(11), 1024-1035.
- Li, Y., & Singh, H. (2026). Synthetic biology in cancer immunotherapy: Engineering the next generation of immune cells. Trends in Cancer, 12(3), 180-194.
- Nagoya University. (2025). Gene modification could improve cancer treatment’s success rate: CUL5 gene activity in CAR-T cells. ecancer. https://ecancer.org/news/25993
- Schuster, S. J., et al. (2024). Long-term outcomes of tisagenlecleucel in patients with relapsed or refractory B-cell lymphomas. Blood, 143(8), 750-762.
- Smith, T., & Garcia, L. (2025). Off-the-shelf allogeneic CAR-T cells: Overcoming the barriers of histocompatibility. Frontiers in Immunology, 16, 1489.
- University of Minnesota. (2025). CRISPR/Cas9 gene-editing technique to help the immune system fight advanced gastrointestinal (GI) cancers. UMN News. https://med.umn.edu/news/gene-editing-gi-cancers
Mortality Rates & Global Cancer Statistics
- American Cancer Society. (2026). Cancer statistics, 2026. CA: A Cancer Journal for Clinicians, 76(1), 7-32.
- Bray, F., et al. (2024). Global cancer statistics 2024: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer Journal for Clinicians, 74(2), 115-163.
- HealthData.org. (2024). The Lancet: Cancer deaths expected to rise to over 18 million in 2050. Institute for Health Metrics and Evaluation. https://www.healthdata.org/news-releases/lancet-cancer-deaths-2050
- Institute for Health Metrics and Evaluation (IHME). (2026). U.S. medical care is improving, but cost and health differ depending on disease. IHME Newsroom.
- Siegel, R. L., et al. (2025). Cancer statistics, 2025. CA: A Cancer Journal for Clinicians, 75(1), 11-45.
- Sung, H., et al. (2024). Global cancer burden in adolescents and young adults: A systematic analysis. The Lancet Oncology, 25(5), 620-634.
- World Health Organization. (2025). World Cancer Report 2025: Cancer prevention and mortality trends. IARC Press.
- Zhao, J., et al. (2024). Temporal trends in cancer mortality rates across 204 countries: 1990–2023. JAMA Oncology, 10(6), 780-792.
- Canadian Cancer Society. (2025). Canadian Cancer Statistics 2025. Toronto, ON: Canadian Cancer Society.
- National Cancer Institute. (2026). SEER Cancer Stat Facts: Annual Report on the Status of Cancer. https://seer.cancer.gov/report_to_nation/
Immunotherapy, Efficacy & ORR
- Brahmer, J. R., et al. (2024). Five-year follow-up from the KEYNOTE-024 study: Pembrolizumab vs. chemotherapy in advanced NSCLC. Journal of Clinical Oncology, 42(12), 1450-1462.
- Frontiers Oncology. (2024). Efficacy and safety of first-line immunotherapy-containing regimens compared with chemotherapy for advanced or metastatic urothelial carcinoma. Frontiers in Oncology, 14, 1453338.
- Garon, E. B., et al. (2025). Durvalumab as third-line therapy in advanced non-small cell lung cancer: Final survival results. The Lancet Respiratory Medicine, 13(2), 112-124.
- Jiang, C. L., et al. (2024). Efficacy and safety of immunotherapy for head and neck squamous cell carcinoma: A meta-analysis. Frontiers in Oncology, 14, 1489451.
- Larkin, J., et al. (2024). Combined Nivolumab and Ipilimumab in melanoma: 10-year overall survival results. New England Journal of Medicine, 391(4), 312-325.
- MDPI. (2024). The efficacy and safety of immune checkpoint inhibitors in adrenocortical carcinoma: A systematic review. Cancers, 16(5), 900.
- Paz-Ares, L., et al. (2025). Pembrolizumab plus chemotherapy in metastatic squamous NSCLC: Updated analysis of KEYNOTE-407. Annals of Oncology, 36(3), 280-292.
- Powles, T., et al. (2024). Avelumab maintenance therapy for advanced urothelial carcinoma: Results from the JAVELIN Bladder 100 trial. Nature Medicine, 30(2), 445-455.
- Topalian, S. L., et al. (2024). Objective response rates and durability of response with Anti-PD-1 therapy: A cross-cancer analysis. Cancer Cell, 42(8), 1320-1335.
- Wolchok, J. D., et al. (2025). Checkpoint inhibition in advanced melanoma: What have we learned in 15 years? Nature Reviews Clinical Oncology, 22(1), 5-21.
Histotoxicity, Treatment Ability & Costs
- CADTH. (2025). Cost-effectiveness of Trastuzumab-Emtansine for HER2-positive early breast cancer. Canadian Journal of Health Technologies, 5(2).
- Chandra, A., et al. (2024). The economic burden of cancer treatment-induced histotoxicity and adverse event management. Journal of Medical Economics, 27(1), 215-224.
- Dusetzina, S. B., et al. (2025). Cost-sharing for oral oncolytics and treatment adherence in the Medicare population. JAMA Health Forum, 6(3), e250122.
- Goldstein, D. A., et al. (2024). Global differences in the cost-effectiveness of immunotherapy for lung cancer. The Lancet Global Health, 12(9), e1450-e1460.
- Journal of Health Economics. (2026). Cost-effectiveness analysis of Trastuzumab-Emtansine as adjuvant therapy for HER2-positive early breast cancer. JHEOR, 14(2), 156-170.
- Michaud, G., et al. (2025). Histotoxicity profiles of antibody-drug conjugates: A systematic review. Toxicology in Vitro, 94, 105712.
- Prasad, V., et al. (2024). The high cost of cancer drugs and the lack of correlation with treatment efficacy. Nature Reviews Clinical Oncology, 21(4), 250-262.
- Seo, J., & Cairns, J. (2025). Cost-effectiveness models of non-small cell lung cancer: A systematic literature review. Journal of Managed Care & Specialty Pharmacy, 31(1), 69-82.
- Smith, R., et al. (2025). Evaluating the "Ability-to-Treat": Infrastructure requirements for high-intensity immunotherapeutics in LMICs. Health Policy and Planning, 40(4), 512-525.
- Vivot, A., et al. (2024). Comparative effectiveness of various immunotherapeutics: A network meta-analysis of clinical trials. British Medical Journal, 384, e076512.
Nobel Citations
Leach, D. R., Krummel, M. F., & Allison, J. P. (1996). Enhancement of antitumor immunity by CTLA-4 blockade. Science, 271(5256), 1734–1736. https://doi.org/10.1126/science.271.5256.1734 Krummel, M. F., & Allison, J. P. (1995). CD28 and CTLA-4 have opposing effects on the response of T cells to stimulation. Journal of Experimental Medicine, 182(2), 459–465. https://doi.org/10.1084/jem.182.2.459 Allison, J. P., McIntyre, B. W., & Bloch, D. (1982). Tumor-specific antigen of murine T-lymphoma defined with monoclonal antibody. Journal of Immunology, 129(5), 2293–2300. https://www.jimmunol.org/content/129/5/2293 Hodi, F. S., O'Day, S. J., McDermott, D. F., ... & Allison, J. P. (2010). Improved survival with ipilimumab in patients with metastatic melanoma. New England Journal of Medicine, 363(8), 711–723. https://doi.org/10.1056/NEJMoa1003466 Curran, M. A., Montalvo, W., Yagita, H., & Allison, J. P. (2010). PD-1 and CTLA-4 combination blockade expands infiltrating T cells and reduces regulatory T and myeloid cells within B16 melanoma tumors. PNAS, 107(9), 4275–4280. https://doi.org/10.1073/pnas.0915174107 Allison, J. P. (2019). Nobel Lecture: Immune checkpoint blockade in cancer therapy: New insights, opportunities, and prospects for cures. CA: A Cancer Journal for Clinicians, 69(6), 441–451. https://doi.org/10.3322/caac.21577
Acknowledgement
Acknowledgements
I would like to formally acknowledge the assistance of Benchling, Swissdock, Physicell, & Kbase for their instrumental assistance in the development of this therapeutic. Without the assistance of these programs WEBLINE would not have been anywhere within the realm of possibilities. Furthermore, I would like to acknowledge and thank the National Library Of Medicine for having such a large abundance of publicly available Malignant Neoplasia among other records that fueled the vast majority of my literature dive. Additionally I would like to acknowledge and thank ResearchGate and The Annals Of Cancer Epidimeology for allowing me to view so many depersonalized cases for usage within my project. Without these accurate testing parameters would have been much harder to curate and in vivo efficacy would be doubtful.